You are browsing environment: FUNGIDB
CAZyme Information: FVEG_03463-t26_1-p1
You are here: Home > Sequence: FVEG_03463-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium verticillioides | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium verticillioides | |||||||||||
| CAZyme ID | FVEG_03463-t26_1-p1 | |||||||||||
| CAZy Family | CBM18|CBM18|CBM18 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.78:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 37 | 313 | 6.1e-86 | 0.9930795847750865 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 226444 | COG3934 | 7.94e-26 | 12 | 325 | 3 | 291 | Endo-1,4-beta-mannosidase [Carbohydrate transport and metabolism]. |
| 395098 | Cellulase | 3.71e-10 | 14 | 316 | 2 | 271 | Cellulase (glycosyl hydrolase family 5). |
| 397119 | Glyco_hydro_2_C | 1.70e-04 | 141 | 298 | 105 | 226 | Glycosyl hydrolases family 2, TIM barrel domain. This family contains beta-galactosidase, beta-mannosidase and beta-glucuronidase activities. |
| 225344 | BglC | 0.002 | 116 | 307 | 133 | 343 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 7.08e-262 | 3 | 366 | 21 | 384 | |
| 5.33e-257 | 2 | 361 | 20 | 379 | |
| 5.33e-257 | 2 | 361 | 20 | 379 | |
| 5.33e-257 | 2 | 361 | 20 | 379 | |
| 5.33e-257 | 2 | 361 | 20 | 379 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.38e-104 | 13 | 347 | 11 | 342 | Crtstal structure of glycoside hydrolase family 5 beta-mannanase from Talaromyces trachyspermus [Talaromyces trachyspermus] |
|
| 3.57e-101 | 13 | 348 | 12 | 344 | Chain A, ENDO-1,4-B-D-MANNANASE [Trichoderma reesei],1QNP_A Chain A, Endo-1,4-b-d-mannanase [Trichoderma reesei],1QNQ_A Chain A, Endo-1,4-b-d-mannanase [Trichoderma reesei],1QNR_A Chain A, Endo-1,4-b-d-mannanase [Trichoderma reesei],1QNS_A Chain A, Endo-1,4-b-d-mannanase [Trichoderma reesei] |
|
| 3.38e-99 | 13 | 349 | 10 | 344 | The ligand-free structure of ManBK from Aspergillus niger BK01 [Aspergillus niger],3WH9_B The ligand-free structure of ManBK from Aspergillus niger BK01 [Aspergillus niger] |
|
| 4.97e-97 | 13 | 348 | 23 | 355 | Chain A, Gh5 Endo-beta-1,4-mannanase [Podospora anserina] |
|
| 2.42e-77 | 11 | 350 | 10 | 359 | Crystal Structure Analysis of the Endo-1,4-beta-mannanase from Rhizomucor miehei [Rhizomucor miehei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.93e-122 | 7 | 352 | 36 | 384 | Mannan endo-1,4-beta-mannosidase B OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=manB PE=1 SV=2 |
|
| 4.61e-109 | 13 | 351 | 40 | 376 | Mannan endo-1,4-beta-mannosidase A OS=Aspergillus aculeatus OX=5053 GN=manA PE=1 SV=1 |
|
| 1.40e-104 | 13 | 348 | 127 | 461 | Probable mannan endo-1,4-beta-mannosidase F OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=manF PE=3 SV=1 |
|
| 2.81e-104 | 13 | 348 | 127 | 461 | Probable mannan endo-1,4-beta-mannosidase F OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=manF PE=3 SV=2 |
|
| 5.91e-104 | 13 | 350 | 51 | 385 | Probable mannan endo-1,4-beta-mannosidase A OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=manA PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
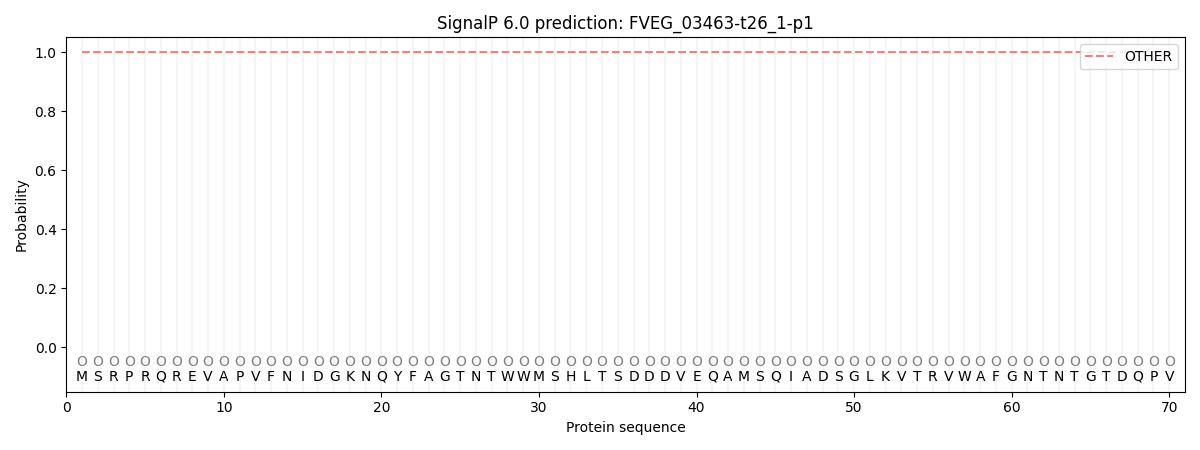
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000051 | 0.000001 |
