You are browsing environment: FUNGIDB
CAZyme Information: FVEG_00488-t26_1-p1
You are here: Home > Sequence: FVEG_00488-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
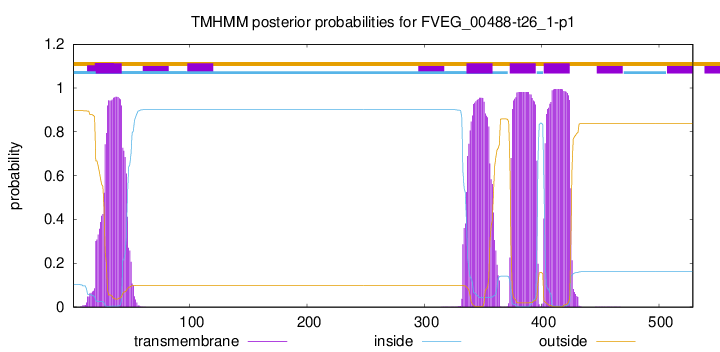
TMHMM annotations
Basic Information help
| Species | Fusarium verticillioides | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium verticillioides | |||||||||||
| CAZyme ID | FVEG_00488-t26_1-p1 | |||||||||||
| CAZy Family | AA16 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT2 | 77 | 312 | 5.7e-26 | 0.9826086956521739 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 133056 | GT2_HAS | 1.23e-113 | 77 | 317 | 1 | 235 | Hyaluronan synthases catalyze polymerization of hyaluronan. Hyaluronan synthases (HASs) are bi-functional glycosyltransferases that catalyze polymerization of hyaluronan. HASs transfer both GlcUA and GlcNAc in beta-(1,3) and beta-(1,4) linkages, respectively to the hyaluronan chain using UDP-GlcNAc and UDP-GlcUA as substrates. HA is made as a free glycan, not attached to a protein or lipid. HASs do not need a primer for HA synthesis; they initiate HA biosynthesis de novo with only UDP-GlcNAc, UDP-GlcUA, and Mg2+. Hyaluronan (HA) is a linear heteropolysaccharide composed of (1-3)-linked beta-D-GlcUA-beta-D-GlcNAc disaccharide repeats. It can be found in vertebrates and a few microbes and is typically on the cell surface or in the extracellular space, but is also found inside mammalian cells. Hyaluronan has several physiochemical and biological functions such as space filling, lubrication, and providing a hydrated matrix through which cells can migrate. |
| 404520 | Glyco_tranf_2_3 | 1.48e-27 | 77 | 313 | 3 | 230 | Glycosyltransferase like family 2. Members of this family of prokaryotic proteins include putative glucosyltransferase, which are involved in bacterial capsule biosynthesis. |
| 133045 | CESA_like | 4.76e-20 | 80 | 252 | 1 | 173 | CESA_like is the cellulose synthase superfamily. The cellulose synthase (CESA) superfamily includes a wide variety of glycosyltransferase family 2 enzymes that share the common characteristic of catalyzing the elongation of polysaccharide chains. The members include cellulose synthase catalytic subunit, chitin synthase, glucan biosynthesis protein and other families of CESA-like proteins. Cellulose synthase catalyzes the polymerization reaction of cellulose, an aggregate of unbranched polymers of beta-1,4-linked glucose residues in plants, most algae, some bacteria and fungi, and even some animals. In bacteria, algae and lower eukaryotes, there is a second unrelated type of cellulose synthase (Type II), which produces acylated cellulose, a derivative of cellulose. Chitin synthase catalyzes the incorporation of GlcNAc from substrate UDP-GlcNAc into chitin, which is a linear homopolymer of beta-(1,4)-linked GlcNAc residues and Glucan Biosynthesis protein catalyzes the elongation of beta-1,2 polyglucose chains of Glucan. |
| 224136 | BcsA | 1.90e-14 | 25 | 431 | 6 | 405 | Glycosyltransferase, catalytic subunit of cellulose synthase and poly-beta-1,6-N-acetylglucosamine synthase [Cell motility]. |
| 395426 | Glycos_transf_2 | 2.02e-05 | 79 | 245 | 1 | 164 | Glycosyl transferase family 2. Diverse family, transferring sugar from UDP-glucose, UDP-N-acetyl- galactosamine, GDP-mannose or CDP-abequose, to a range of substrates including cellulose, dolichol phosphate and teichoic acids. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 529 | 1 | 529 | |
| 0.0 | 1 | 529 | 1 | 529 | |
| 0.0 | 1 | 529 | 1 | 529 | |
| 0.0 | 1 | 529 | 1 | 529 | |
| 0.0 | 1 | 529 | 1 | 529 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 1 | 529 | 1 | 526 | Type 2 glycosyltransferase OS=Gibberella zeae (strain ATCC MYA-4620 / CBS 123657 / FGSC 9075 / NRRL 31084 / PH-1) OX=229533 GN=GT2 PE=2 SV=1 |
|
| 7.47e-233 | 37 | 467 | 32 | 461 | Type 2 glycosyltransferase OS=Magnaporthe oryzae (strain 70-15 / ATCC MYA-4617 / FGSC 8958) OX=242507 GN=GT2 PE=1 SV=1 |
|
| 8.79e-144 | 37 | 463 | 7 | 429 | Type 2 glycosyltransferase OS=Zymoseptoria tritici (strain CBS 115943 / IPO323) OX=336722 GN=GT2 PE=3 SV=1 |
|
| 4.25e-10 | 156 | 433 | 211 | 468 | Hyaluronan synthase 3 OS=Homo sapiens OX=9606 GN=HAS3 PE=1 SV=3 |
|
| 5.63e-10 | 156 | 433 | 212 | 469 | Hyaluronan synthase 3 OS=Mus musculus OX=10090 GN=Has3 PE=1 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999071 | 0.000952 |