You are browsing environment: FUNGIDB
CAZyme Information: FPRO_05401-t41_1-p1
You are here: Home > Sequence: FPRO_05401-t41_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium proliferatum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium proliferatum | |||||||||||
| CAZyme ID | FPRO_05401-t41_1-p1 | |||||||||||
| CAZy Family | CE12 | |||||||||||
| CAZyme Description | related to beta-glucosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.21:2 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 32 | 243 | 1.8e-61 | 0.9814814814814815 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 224389 | BglX | 2.37e-68 | 25 | 366 | 48 | 382 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| 396478 | Glyco_hydro_3_C | 6.45e-46 | 530 | 695 | 53 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
| 185053 | PRK15098 | 4.77e-42 | 61 | 805 | 118 | 736 | beta-glucosidase BglX. |
| 395747 | Glyco_hydro_3 | 1.36e-35 | 61 | 278 | 86 | 316 | Glycosyl hydrolase family 3 N terminal domain. |
| 178629 | PLN03080 | 5.34e-29 | 56 | 729 | 104 | 648 | Probable beta-xylosidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 846 | 1 | 846 | |
| 0.0 | 1 | 846 | 1 | 861 | |
| 0.0 | 1 | 846 | 1 | 861 | |
| 0.0 | 1 | 846 | 1 | 846 | |
| 0.0 | 1 | 845 | 1 | 847 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.04e-146 | 6 | 816 | 7 | 817 | Crystal structure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus],3AC0_B Crystal structure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus],3AC0_C Crystal structure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus],3AC0_D Crystal structure of Beta-glucosidase from Kluyveromyces marxianus in complex with glucose [Kluyveromyces marxianus] |
|
| 4.15e-142 | 6 | 816 | 7 | 817 | Crystal structure of Se-Met labeled Beta-glucosidase from Kluyveromyces marxianus [Kluyveromyces marxianus],3ABZ_B Crystal structure of Se-Met labeled Beta-glucosidase from Kluyveromyces marxianus [Kluyveromyces marxianus],3ABZ_C Crystal structure of Se-Met labeled Beta-glucosidase from Kluyveromyces marxianus [Kluyveromyces marxianus],3ABZ_D Crystal structure of Se-Met labeled Beta-glucosidase from Kluyveromyces marxianus [Kluyveromyces marxianus] |
|
| 6.93e-105 | 6 | 816 | 6 | 649 | Chain A, Thermostable beta-glucosidase B [Acetivibrio thermocellus AD2],7MS2_B Chain B, Thermostable beta-glucosidase B [Acetivibrio thermocellus AD2] |
|
| 8.23e-91 | 6 | 821 | 5 | 699 | Structure of beta-glucosidase 3B from Thermotoga neapolitana in complex with glycerol [Thermotoga neapolitana DSM 4359],2X41_A Structure of beta-glucosidase 3B from Thermotoga neapolitana in complex with glucose [Thermotoga neapolitana DSM 4359] |
|
| 1.15e-89 | 6 | 821 | 5 | 699 | Structure of beta-glucosidase 3B from Thermotoga neapolitana in complex with alpha-D-glucose [Thermotoga neapolitana DSM 4359] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.21e-151 | 10 | 817 | 12 | 802 | Probable beta-glucosidase H OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=bglH PE=3 SV=1 |
|
| 5.15e-150 | 4 | 817 | 7 | 802 | Probable beta-glucosidase H OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=bglH PE=3 SV=1 |
|
| 1.02e-149 | 10 | 817 | 12 | 802 | Probable beta-glucosidase H OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=bglH PE=3 SV=1 |
|
| 5.60e-149 | 10 | 817 | 12 | 802 | Probable beta-glucosidase H OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=bglH PE=3 SV=1 |
|
| 2.05e-148 | 1 | 822 | 1 | 842 | Probable beta-glucosidase J OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=bglJ PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
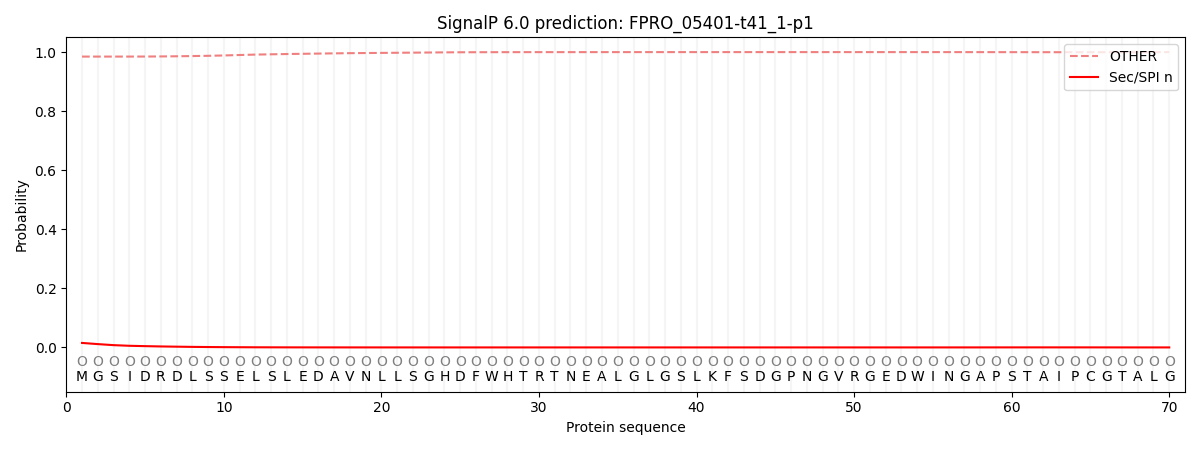
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.985952 | 0.014062 |
