You are browsing environment: FUNGIDB
CAZyme Information: FPRN_04567-t42_1-p1
You are here: Home > Sequence: FPRN_04567-t42_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium proliferatum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium proliferatum | |||||||||||
| CAZyme ID | FPRN_04567-t42_1-p1 | |||||||||||
| CAZy Family | CBM67|GH78 | |||||||||||
| CAZyme Description | related to protein kinase C pathway protein SMP3 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT22 | 5 | 397 | 1.1e-78 | 0.9897172236503856 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 281842 | Glyco_transf_22 | 1.62e-44 | 5 | 371 | 2 | 372 | Alg9-like mannosyltransferase family. Members of this family are mannosyltransferase enzymes. At least some members are localized in endoplasmic reticulum and involved in GPI anchor biosynthesis. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 517 | 1 | 517 | |
| 0.0 | 1 | 517 | 1 | 517 | |
| 0.0 | 1 | 517 | 1 | 517 | |
| 0.0 | 1 | 517 | 1 | 516 | |
| 0.0 | 1 | 517 | 1 | 517 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 1 | 517 | 1 | 517 | GPI mannosyltransferase 4 OS=Gibberella zeae (strain ATCC MYA-4620 / CBS 123657 / FGSC 9075 / NRRL 31084 / PH-1) OX=229533 GN=SMP3 PE=3 SV=2 |
|
| 1.20e-180 | 1 | 514 | 1 | 544 | GPI mannosyltransferase 4 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=smp3 PE=3 SV=2 |
|
| 4.23e-180 | 1 | 517 | 1 | 543 | GPI mannosyltransferase 4 OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=smp3 PE=3 SV=1 |
|
| 3.84e-166 | 1 | 509 | 1 | 534 | GPI mannosyltransferase 4 OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=smp3 PE=3 SV=2 |
|
| 6.96e-100 | 10 | 487 | 13 | 495 | GPI mannosyltransferase 4 OS=Yarrowia lipolytica (strain CLIB 122 / E 150) OX=284591 GN=SMP3 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.065658 | 0.934308 | CS pos: 23-24. Pr: 0.7640 |
TMHMM Annotations download full data without filtering help
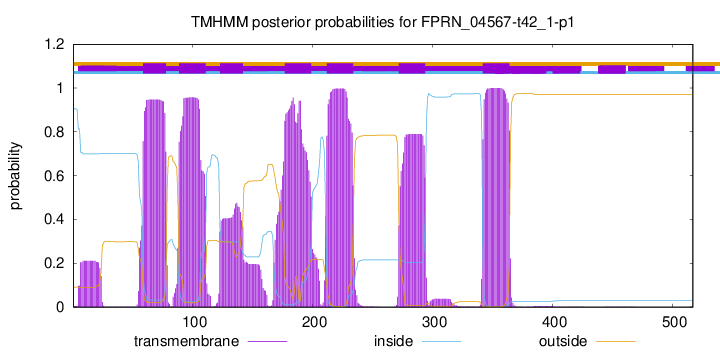
| Start | End |
|---|---|
| 59 | 78 |
| 89 | 111 |
| 123 | 142 |
| 177 | 199 |
| 212 | 234 |
| 272 | 294 |
| 342 | 364 |
