You are browsing environment: FUNGIDB
CAZyme Information: FGRAMPH1_01T25365-p1
You are here: Home > Sequence: FGRAMPH1_01T25365-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium graminearum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium graminearum | |||||||||||
| CAZyme ID | FGRAMPH1_01T25365-p1 | |||||||||||
| CAZy Family | GT2|GT2 | |||||||||||
| CAZyme Description | laccase precursor | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA1 | 136 | 631 | 2.7e-113 | 0.9804469273743017 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 274555 | ascorbase | 1.30e-69 | 125 | 631 | 1 | 517 | L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. |
| 259953 | CuRO_2_MCO_like_1 | 7.74e-68 | 260 | 421 | 1 | 163 | The second cupredoxin domain of uncharacterized multicopper oxidase. Multicopper Oxidases (MCOs) are multi-domain enzymes that are able to couple oxidation of substrates with reduction of dioxygen to water. MCOs oxidise their substrate by accepting electrons at a mononuclear copper centre and transferring them to a trinuclear copper centre which binds a dioxygen. The dioxygen, following the transfer of four electrons, is reduced to two molecules of water. These MCOs are capable of oxidizing a vast range of substrates, varying from aromatic to inorganic compounds such as metals. This family of MCOs is composed of three cupredoxin domains that include one mononuclear and one trinuclear copper center. The copper ions are bound in several sites: Type 1, Type 2, and/or Type 3. The ensemble of types 2 and 3 copper is called a trinuclear cluster. MCOs oxidize their substrate by accepting electrons at a mononuclear copper center and transferring them to the active site trinuclear copper center. The cupredoxin domain 2 of 3-domain MCOs has lost the ability to bind copper. |
| 215324 | PLN02604 | 2.09e-67 | 125 | 631 | 24 | 540 | oxidoreductase |
| 259926 | CuRO_1_Diphenol_Ox | 7.07e-67 | 126 | 244 | 1 | 119 | The first cupredoxin domain of fungal laccase, diphenol oxidase. Diphenol oxidase belongs to the laccase family. It catalyzes the initial steps in melanin biosynthesis from diphenols. Melanin is one of the virulence factors of infectious fungi. In the pathogenesis of C. neoformans, melanin pigments have been shown to protect the fungal cells from oxidative and microbicidal activities of host defense systems. Laccase is a blue multicopper oxidase (MCO) which catalyzes the oxidation of a variety aromatic - notably phenolic and inorganic substances coupled to the reduction of molecular oxygen to water. It has been implicated in a wide spectrum of biological activities and, in particular, plays a key role in morphogenesis, development and lignin metabolism. Although MCOs have diverse functions, majority of them have three cupredoxin domain repeats that include one mononuclear and one trinuclear copper center. The copper ions are bound in several sites: Type 1, Type 2, and/or Type 3. The ensemble of types 2 and 3 copper is called a trinuclear cluster. MCOs oxidize their substrate by accepting electrons at a mononuclear copper center and transferring them to the active site trinuclear copper center. The cupredoxin domain 1 of 3-domain MCOs contains part the trinuclear copper binding site, which is located at the interface of domains 1 and 3. |
| 177843 | PLN02191 | 2.73e-63 | 121 | 631 | 19 | 540 | L-ascorbate oxidase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 677 | 1 | 677 | |
| 0.0 | 1 | 677 | 1 | 677 | |
| 0.0 | 1 | 677 | 1 | 677 | |
| 0.0 | 1 | 677 | 1 | 677 | |
| 0.0 | 1 | 675 | 1 | 672 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.26e-75 | 110 | 636 | 51 | 541 | Structure of the L499M mutant of the laccase from B.aclada [Botrytis aclada] |
|
| 3.40e-75 | 110 | 636 | 51 | 541 | Crystal structure of laccase from Botrytis aclada at 1.67 A resolution [Botrytis aclada],4X4K_A Structure of laccase from Botrytis aclada with full copper content [Botrytis aclada] |
|
| 9.50e-68 | 130 | 644 | 7 | 486 | Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_B Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_C Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_D Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_E Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_F Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae] |
|
| 1.86e-67 | 119 | 638 | 14 | 505 | Crystal structure of a laccase-like multicopper oxidase McoG from from Aspergillus niger [Aspergillus niger] |
|
| 1.90e-67 | 119 | 638 | 15 | 506 | Crystal structure of a laccase-like multicopper oxidase McoG from Aspergillus niger bound to zinc [Aspergillus niger] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.17e-99 | 85 | 655 | 21 | 578 | Laccase-1 OS=Cryptococcus neoformans var. grubii serotype A (strain H99 / ATCC 208821 / CBS 10515 / FGSC 9487) OX=235443 GN=LAC1 PE=1 SV=1 |
|
| 3.39e-98 | 99 | 655 | 35 | 578 | Laccase-1 OS=Cryptococcus neoformans var. neoformans serotype D (strain B-3501A) OX=283643 GN=LAC1 PE=1 SV=1 |
|
| 1.88e-95 | 99 | 655 | 35 | 578 | Laccase-2 OS=Cryptococcus neoformans var. grubii serotype A (strain H99 / ATCC 208821 / CBS 10515 / FGSC 9487) OX=235443 GN=LAC2 PE=3 SV=2 |
|
| 7.77e-77 | 125 | 662 | 73 | 572 | Oxidoreductase OpS5 OS=Beauveria bassiana (strain ARSEF 2860) OX=655819 GN=OpS5 PE=1 SV=1 |
|
| 5.21e-75 | 126 | 655 | 65 | 566 | Oxydoreductase ptaK OS=Pestalotiopsis fici (strain W106-1 / CGMCC3.15140) OX=1229662 GN=ptaK PE=2 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
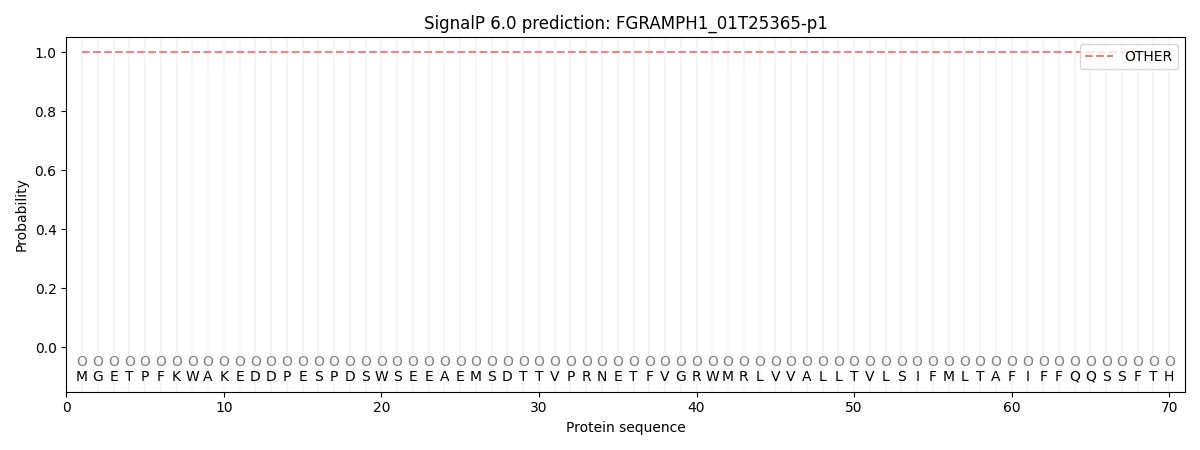
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999975 | 0.000020 |

