You are browsing environment: FUNGIDB
CAZyme Information: FGRAMPH1_01T20781-p1
You are here: Home > Sequence: FGRAMPH1_01T20781-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium graminearum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium graminearum | |||||||||||
| CAZyme ID | FGRAMPH1_01T20781-p1 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | udp-glucosyltransferase yojk | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT1 | 21 | 407 | 1.3e-40 | 0.9057591623036649 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 340817 | GT1_Gtf-like | 2.06e-37 | 1 | 406 | 17 | 404 | UDP-glycosyltransferases and similar proteins. This family includes the Gtfs, a group of homologous glycosyltransferases involved in the final stages of the biosynthesis of antibiotics vancomycin and related chloroeremomycin. Gtfs transfer sugar moieties from an activated NDP-sugar donor to the oxidatively cross-linked heptapeptide core of vancomycin group antibiotics. The core structure is important for the bioactivity of the antibiotics. |
| 224732 | YjiC | 2.00e-29 | 3 | 413 | 20 | 405 | UDP:flavonoid glycosyltransferase YjiC, YdhE family [Carbohydrate transport and metabolism]. |
| 273616 | MGT | 1.14e-27 | 3 | 406 | 14 | 389 | glycosyltransferase, MGT family. This model describes the MGT (macroside glycosyltransferase) subfamily of the UDP-glucuronosyltransferase family. Members include a number of glucosyl transferases for macrolide antibiotic inactivation, but also include transferases of glucose-related sugars for macrolide antibiotic production. [Cellular processes, Toxin production and resistance] |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.21e-303 | 1 | 413 | 1 | 413 | |
| 9.02e-302 | 1 | 413 | 17 | 429 | |
| 4.71e-300 | 1 | 413 | 1 | 413 | |
| 3.40e-293 | 1 | 411 | 1 | 411 | |
| 4.59e-244 | 1 | 412 | 17 | 427 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.55e-14 | 4 | 407 | 32 | 420 | The crystal structure of macrolide glycosyltransferases: A blueprint for antibiotic engineering [Streptomyces antibioticus],2IYA_B The crystal structure of macrolide glycosyltransferases: A blueprint for antibiotic engineering [Streptomyces antibioticus] |
|
| 3.64e-12 | 4 | 407 | 40 | 412 | Crystal Structure of CalG2, Calicheamicin Glycosyltransferase, TDP and calicheamicin T0 bound form [Micromonospora echinospora],3RSC_B Crystal Structure of CalG2, Calicheamicin Glycosyltransferase, TDP and calicheamicin T0 bound form [Micromonospora echinospora] |
|
| 3.65e-12 | 4 | 407 | 40 | 412 | Crystal Structure of CalG2, Calicheamicin Glycosyltransferase, TDP bound form [Micromonospora echinospora],3IAA_B Crystal Structure of CalG2, Calicheamicin Glycosyltransferase, TDP bound form [Micromonospora echinospora] |
|
| 1.15e-10 | 82 | 393 | 122 | 399 | Crystal structure of glycosyltransferase VinC involved in the biosynthesis of vicenistatin [Streptomyces halstedii],3WAD_B Crystal structure of glycosyltransferase VinC involved in the biosynthesis of vicenistatin [Streptomyces halstedii],3WAG_A Crystal structure of glycosyltransferase VinC in complex with DTDP [Streptomyces halstedii],3WAG_B Crystal structure of glycosyltransferase VinC in complex with DTDP [Streptomyces halstedii] |
|
| 8.41e-10 | 234 | 405 | 225 | 396 | The crystal structure of macrolide glycosyltransferases: A blueprint for antibiotic engineering [Streptomyces antibioticus],2IYF_B The crystal structure of macrolide glycosyltransferases: A blueprint for antibiotic engineering [Streptomyces antibioticus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.67e-41 | 1 | 407 | 19 | 421 | 4'-demethylrebeccamycin synthase OS=Lentzea aerocolonigenes OX=68170 GN=rebG PE=1 SV=1 |
|
| 2.26e-31 | 3 | 407 | 26 | 435 | UDP-glucosyltransferase A1 OS=Starmerella bombicola OX=75736 GN=ugtA1 PE=1 SV=1 |
|
| 1.99e-27 | 3 | 411 | 24 | 431 | UDP-glucosyltransferase B1 OS=Starmerella bombicola OX=75736 GN=ugtB1 PE=1 SV=1 |
|
| 4.48e-24 | 4 | 397 | 38 | 448 | UDP-glucosyltransferase 1 OS=Beauveria bassiana (strain ARSEF 2860) OX=655819 GN=BBA_08686 PE=2 SV=1 |
|
| 4.50e-09 | 234 | 405 | 225 | 396 | Oleandomycin glycosyltransferase OS=Streptomyces antibioticus OX=1890 GN=oleD PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
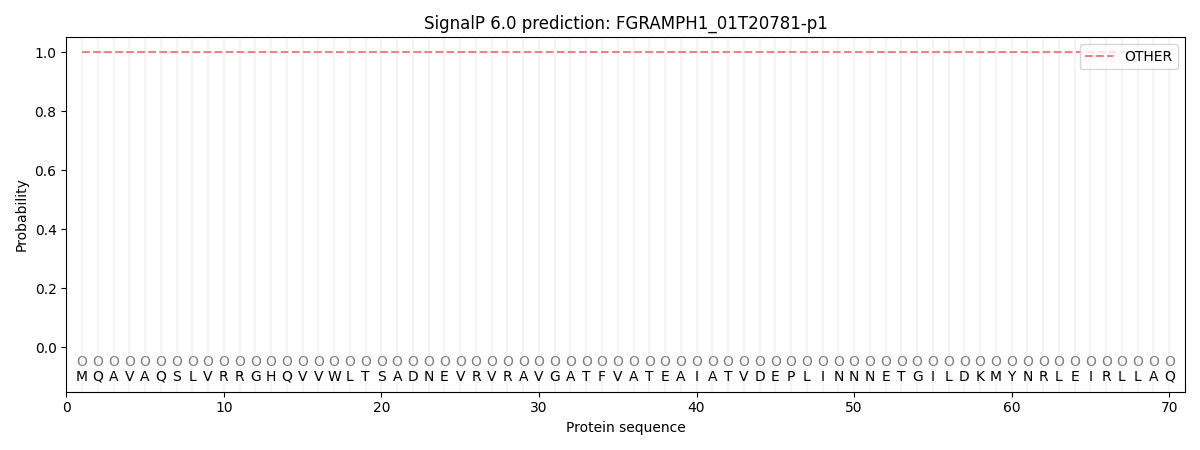
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000083 | 0.000000 |
