You are browsing environment: FUNGIDB
CAZyme Information: FGRAMPH1_01T16515-p1
You are here: Home > Sequence: FGRAMPH1_01T16515-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium graminearum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium graminearum | |||||||||||
| CAZyme ID | FGRAMPH1_01T16515-p1 | |||||||||||
| CAZy Family | GH31 | |||||||||||
| CAZyme Description | pectate lyase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL3 | 30 | 215 | 3.5e-80 | 0.9896907216494846 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397360 | Pectate_lyase | 2.25e-117 | 23 | 220 | 1 | 200 | Pectate lyase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.53e-177 | 1 | 231 | 1 | 231 | |
| 1.53e-177 | 1 | 231 | 1 | 231 | |
| 8.83e-177 | 1 | 231 | 1 | 231 | |
| 3.60e-176 | 1 | 231 | 1 | 231 | |
| 7.22e-167 | 1 | 231 | 1 | 231 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.97e-29 | 41 | 164 | 11 | 135 | Crystal Structure Of Pectate Lyase From Bacillus Sp. Strain Ksm-P15. [Bacillus sp. KSM-P15] |
|
| 8.75e-22 | 29 | 165 | 3 | 139 | The liganded structure of C. bescii family 3 pectate lyase [Caldicellulosiruptor bescii DSM 6725],4EW9_B The liganded structure of C. bescii family 3 pectate lyase [Caldicellulosiruptor bescii DSM 6725] |
|
| 8.95e-22 | 29 | 165 | 4 | 140 | The crystal structure of family 3 pectate lyase from Caldicellulosiruptor bescii [Caldicellulosiruptor bescii],3T9G_B The crystal structure of family 3 pectate lyase from Caldicellulosiruptor bescii [Caldicellulosiruptor bescii] |
|
| 7.99e-21 | 29 | 165 | 12 | 148 | C. bescii Family 3 pectate lyase double mutant K108A in complex with trigalacturonic acid [Caldicellulosiruptor bescii DSM 6725],4Z03_B C. bescii Family 3 pectate lyase double mutant K108A in complex with trigalacturonic acid [Caldicellulosiruptor bescii DSM 6725] |
|
| 7.99e-21 | 29 | 165 | 12 | 148 | C. bescii Family 3 pectate lyase mutant E84A [Caldicellulosiruptor bescii DSM 6725],4Z05_B C. bescii Family 3 pectate lyase mutant E84A [Caldicellulosiruptor bescii DSM 6725] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.33e-96 | 3 | 231 | 2 | 232 | Probable pectate lyase F OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=plyF PE=3 SV=1 |
|
| 3.68e-96 | 13 | 231 | 12 | 231 | Probable pectate lyase F OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=plyF PE=3 SV=2 |
|
| 4.41e-95 | 3 | 231 | 2 | 232 | Probable pectate lyase F OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=plyF PE=3 SV=1 |
|
| 8.89e-95 | 3 | 231 | 2 | 232 | Probable pectate lyase F OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=plyF PE=3 SV=1 |
|
| 2.07e-93 | 3 | 231 | 2 | 232 | Probable pectate lyase F OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=plyF PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
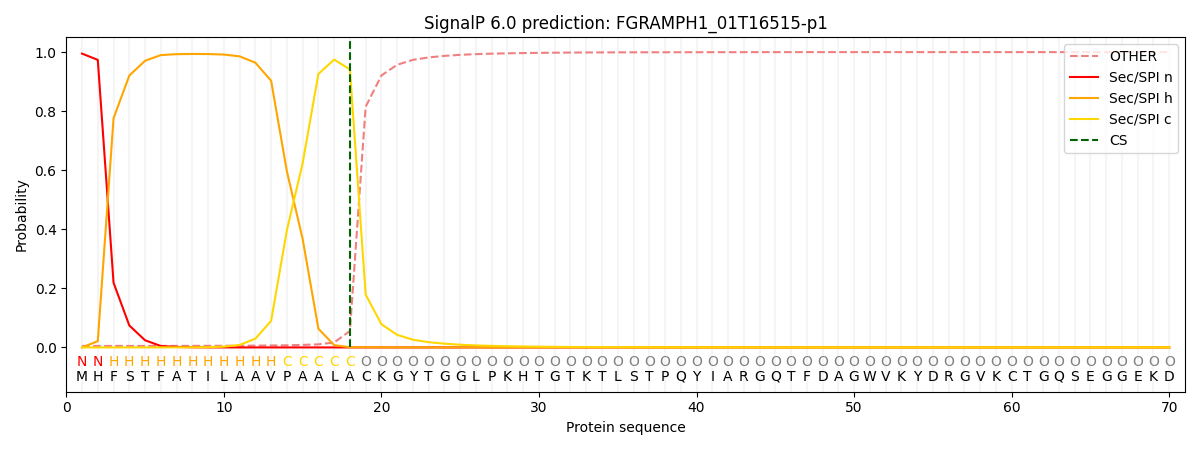
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.007131 | 0.992829 | CS pos: 18-19. Pr: 0.9415 |
