You are browsing environment: FUNGIDB
CAZyme Information: FGRAMPH1_01T16301-p1
You are here: Home > Sequence: FGRAMPH1_01T16301-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium graminearum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium graminearum | |||||||||||
| CAZyme ID | FGRAMPH1_01T16301-p1 | |||||||||||
| CAZy Family | GH31 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| PL42 | 56 | 281 | 6.3e-28 | 0.722972972972973 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 406346 | BNR_4 | 8.29e-29 | 71 | 338 | 8 | 272 | BNR repeat-containing family member. BNR_4 is a family which carries the unique sequence motif SxDxGxTW which is so characteristic of the repeats of the BNR family, pfam02012. It is unclear whether or not this unit is repeated throughout the sequences of this family, but if it is then the family is likely to be bacterial neuraminidase. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 8 | 445 | 1 | 438 | |
| 3.60e-314 | 8 | 445 | 1 | 438 | |
| 5.11e-314 | 8 | 445 | 1 | 438 | |
| 1.11e-294 | 8 | 445 | 1 | 437 | |
| 5.67e-279 | 8 | 444 | 1 | 437 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.47e-13 | 109 | 338 | 90 | 313 | Glycoside Hydrolase BT3686 [Bacteroides thetaiotaomicron VPI-5482],5MUL_A Glycoside Hydrolase BT3686 bound to Glucuronic Acid [Bacteroides thetaiotaomicron VPI-5482] |
|
| 2.59e-12 | 109 | 338 | 95 | 318 | Glycoside Hydrolase BACINT_00347 [Bacteroides intestinalis DSM 17393] |
|
| 1.08e-11 | 109 | 297 | 90 | 281 | Glycoside Hydrolase BACCELL_00856 [Bacteroides cellulosilyticus DSM 14838] |
|
| 1.81e-11 | 109 | 338 | 73 | 296 | Crystal structure of a putative neuraminidase (BACOVA_03493) from Bacteroides ovatus ATCC 8483 at 1.74 A resolution [Bacteroides ovatus ATCC 8483] |
|
| 1.79e-08 | 51 | 253 | 33 | 238 | Chain A, L-Rhamnose-alpha-1,4-D-glucuronate lyase [Fusarium oxysporum],7ESM_A Chain A, L-rhamnose-alpha-1,4-D-glucuronate lyase [Fusarium oxysporum] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
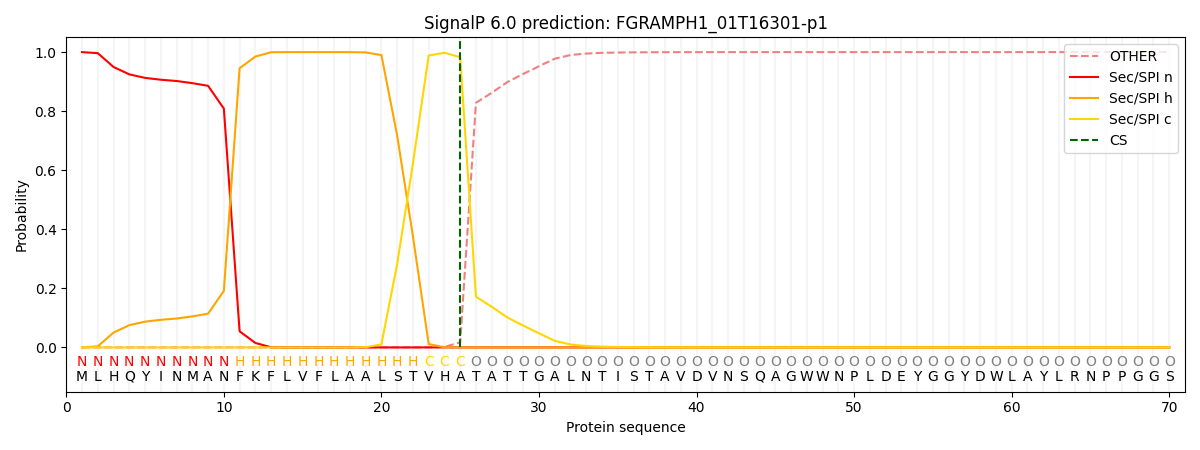
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000214 | 0.999773 | CS pos: 25-26. Pr: 0.9827 |

