You are browsing environment: FUNGIDB
CAZyme Information: FGRAMPH1_01T07723-p1
You are here: Home > Sequence: FGRAMPH1_01T07723-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium graminearum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium graminearum | |||||||||||
| CAZyme ID | FGRAMPH1_01T07723-p1 | |||||||||||
| CAZy Family | CBM508 | |||||||||||
| CAZyme Description | glycosyltransferase family 8 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT8 | 15 | 285 | 4.1e-40 | 0.8832684824902723 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 133018 | GT8_Glycogenin | 3.65e-65 | 16 | 304 | 2 | 236 | Glycogenin belongs the GT 8 family and initiates the biosynthesis of glycogen. Glycogenin initiates the biosynthesis of glycogen by incorporating glucose residues through a self-glucosylation reaction at a Tyr residue, and then acts as substrate for chain elongation by glycogen synthase and branching enzyme. It contains a conserved DxD motif and an N-terminal beta-alpha-beta Rossmann-like fold that are common to the nucleotide-binding domains of most glycosyltransferases. The DxD motif is essential for coordination of the catalytic divalent cation, most commonly Mn2+. Glycogenin can be classified as a retaining glycosyltransferase, based on the relative anomeric stereochemistry of the substrate and product in the reaction catalyzed. It is placed in glycosyltransferase family 8 which includes lipopolysaccharide glucose and galactose transferases and galactinol synthases. |
| 215090 | PLN00176 | 1.20e-16 | 18 | 301 | 27 | 288 | galactinol synthase |
| 133037 | GT8_A4GalT_like | 7.55e-13 | 25 | 282 | 10 | 246 | A4GalT_like proteins catalyze the addition of galactose or glucose residues to the lipooligosaccharide (LOS) or lipopolysaccharide (LPS) of the bacterial cell surface. The members of this family of glycosyltransferases catalyze the addition of galactose or glucose residues to the lipooligosaccharide (LOS) or lipopolysaccharide (LPS) of the bacterial cell surface. The enzymes exhibit broad substrate specificities. The known functions found in this family include: Alpha-1,4-galactosyltransferase, LOS-alpha-1,3-D-galactosyltransferase, UDP-glucose:(galactosyl) LPS alpha1,2-glucosyltransferase, UDP-galactose: (glucosyl) LPS alpha1,2-galactosyltransferase, and UDP-glucose:(glucosyl) LPS alpha1,2-glucosyltransferase. Alpha-1,4-galactosyltransferase from N. meningitidis adds an alpha-galactose from UDP-Gal (the donor) to a terminal lactose (the acceptor) of the LOS structure of outer membrane. LOSs are virulence factors that enable the organism to evade the immune system of host cells. In E. coli, the three alpha-1,2-glycosyltransferases, that are involved in the synthesis of the outer core region of the LPS, are all members of this family. The three enzymes share 40 % of sequence identity, but have different sugar donor or acceptor specificities, representing the structural diversity of LPS. |
| 279798 | Glyco_transf_8 | 9.27e-08 | 23 | 285 | 7 | 251 | Glycosyl transferase family 8. This family includes enzymes that transfer sugar residues to donor molecules. Members of this family are involved in lipopolysaccharide biosynthesis and glycogen synthesis. This family includes Lipopolysaccharide galactosyltransferase, lipopolysaccharide glucosyltransferase 1, and glycogenin glucosyltransferase. |
| 227884 | Gnt1 | 2.89e-07 | 70 | 254 | 138 | 320 | Alpha-N-acetylglucosamine transferase [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.47e-249 | 1 | 322 | 1 | 322 | |
| 2.47e-249 | 1 | 322 | 1 | 322 | |
| 5.14e-242 | 1 | 322 | 1 | 322 | |
| 2.44e-240 | 1 | 322 | 1 | 322 | |
| 2.25e-236 | 1 | 322 | 1 | 322 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.02e-13 | 17 | 290 | 10 | 247 | UDP-galactose:glucoside-Skp1 alpha-D-galactosyltransferase with bound UDP and Platinum [Globisporangium ultimum],6MW8_A UDP-galactose:glucoside-Skp1 alpha-D-galactosyltransferase with bound UDP and Manganese [Globisporangium ultimum] |
|
| 5.14e-06 | 15 | 298 | 5 | 237 | Crystal Structure of Human Glycogenin-1 (GYG1) T83M mutant complexed with manganese and UDP [Homo sapiens],3RMW_A Crystal Structure of Human Glycogenin-1 (GYG1) T83M mutant complexed with manganese and UDP-glucose [Homo sapiens] |
|
| 5.14e-06 | 15 | 298 | 5 | 237 | Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and UDP, in a triclinic closed form [Homo sapiens],3T7M_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and UDP, in a triclinic closed form [Homo sapiens],3T7N_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and UDP, in a monoclinic closed form [Homo sapiens],3T7N_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and UDP, in a monoclinic closed form [Homo sapiens],3T7O_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP-Glucose and glucose [Homo sapiens],3T7O_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP-Glucose and glucose [Homo sapiens],3U2U_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and maltotetraose [Homo sapiens],3U2U_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and maltotetraose [Homo sapiens],3U2V_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and maltohexaose [Homo sapiens],3U2V_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and maltohexaose [Homo sapiens],3U2X_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and 1'-deoxyglucose [Homo sapiens],3U2X_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese, UDP and 1'-deoxyglucose [Homo sapiens] |
|
| 5.71e-06 | 15 | 298 | 26 | 258 | Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese [Homo sapiens] |
|
| 6.88e-06 | 15 | 298 | 5 | 237 | Crystal Structure of Human Glycogenin-1 (GYG1), apo form [Homo sapiens],3QVB_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and UDP [Homo sapiens],3U2W_A Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and glucose or a glucal species [Homo sapiens],3U2W_B Crystal Structure of Human Glycogenin-1 (GYG1) complexed with manganese and glucose or a glucal species [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.25e-17 | 18 | 301 | 27 | 289 | Galactinol synthase 4 OS=Arabidopsis thaliana OX=3702 GN=GOLS4 PE=2 SV=1 |
|
| 1.55e-16 | 18 | 301 | 28 | 289 | Galactinol synthase 2 OS=Solanum lycopersicum OX=4081 GN=GOLS2 PE=2 SV=1 |
|
| 2.02e-16 | 18 | 301 | 27 | 288 | Galactinol synthase 1 OS=Ajuga reptans OX=38596 GN=GOLS1 PE=1 SV=1 |
|
| 2.36e-16 | 18 | 301 | 24 | 275 | Galactinol synthase 1 OS=Solanum lycopersicum OX=4081 GN=GOLS1 PE=2 SV=1 |
|
| 7.65e-16 | 10 | 301 | 25 | 296 | Galactinol synthase 1 OS=Arabidopsis thaliana OX=3702 GN=GOLS1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
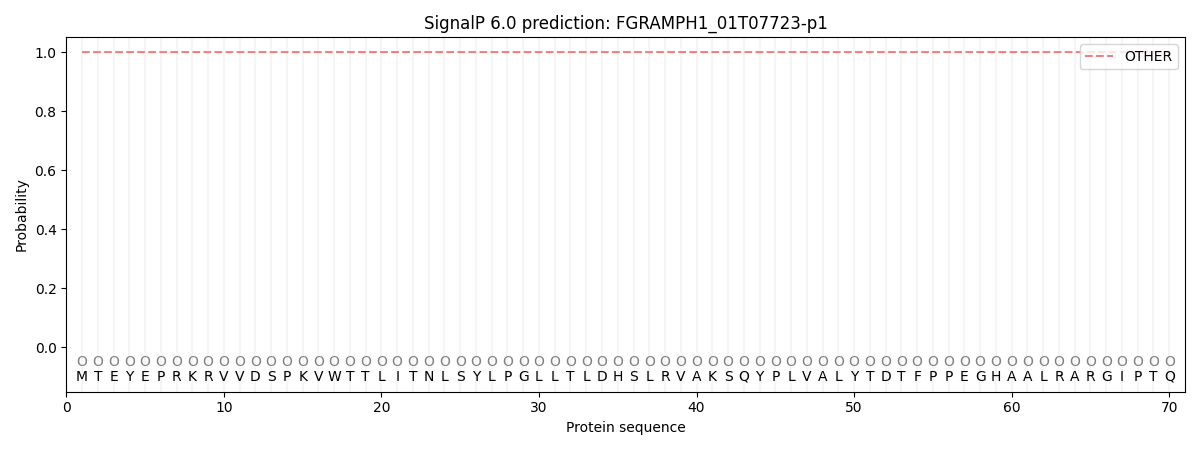
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000060 | 0.000000 |
