You are browsing environment: FUNGIDB
CAZyme Information: EPrPVT00000021297-p1
You are here: Home > Sequence: EPrPVT00000021297-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Phytopythium vexans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Pythiaceae; Phytopythium; Phytopythium vexans | |||||||||||
| CAZyme ID | EPrPVT00000021297-p1 | |||||||||||
| CAZy Family | GH72 | |||||||||||
| CAZyme Description | Thioredoxin reductase 1 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA17 | 9 | 215 | 1.5e-44 | 0.9387755102040817 |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 6.29e-27 | 15 | 162 | 14 | 190 | |
| 7.35e-27 | 14 | 159 | 15 | 181 | |
| 1.37e-26 | 26 | 168 | 48 | 205 | |
| 1.70e-26 | 8 | 175 | 70 | 265 | |
| 2.57e-26 | 15 | 162 | 78 | 254 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.47e-21 | 64 | 154 | 67 | 168 | Chain A, Lytic Polysaccharide Monooxygenase [Phytophthora infestans T30-4],6Z5Y_B Chain B, Lytic Polysaccharide Monooxygenase [Phytophthora infestans T30-4] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
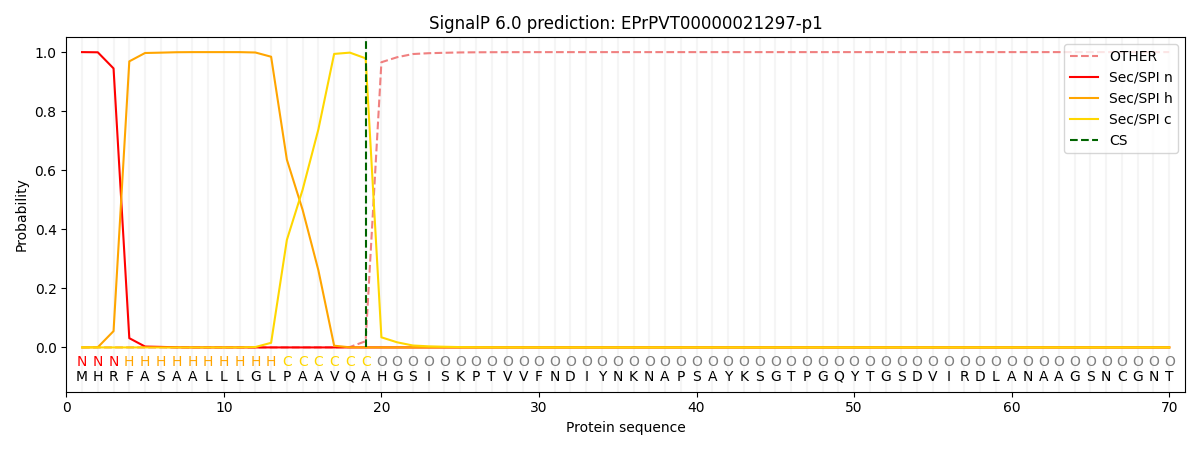
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000225 | 0.999736 | CS pos: 19-20. Pr: 0.9789 |
