You are browsing environment: FUNGIDB
CAZyme Information: EPrPRT00000013096-p1
You are here: Home > Sequence: EPrPRT00000013096-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Pythium arrhenomanes | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Pythiaceae; Pythium; Pythium arrhenomanes | |||||||||||
| CAZyme ID | EPrPRT00000013096-p1 | |||||||||||
| CAZy Family | AA1 | |||||||||||
| CAZyme Description | Alpha mannosyltransferase. | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.122:14 | - |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT10 | 599 | 759 | 5.9e-35 | 0.42939481268011526 |
| GT31 | 83 | 287 | 2.6e-32 | 0.96875 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 250845 | Galactosyl_T | 6.10e-18 | 81 | 291 | 1 | 196 | Galactosyltransferase. This family includes the galactosyltransferases UDP-galactose:2-acetamido-2-deoxy-D-glucose3beta-galactosyltransferase and UDP-Gal:beta-GlcNAc beta 1,3-galactosyltranferase. Specific galactosyltransferases transfer galactose to GlcNAc terminal chains in the synthesis of the lacto-series oligosaccharides types 1 and 2. |
| 395683 | Glyco_transf_10 | 7.29e-15 | 620 | 781 | 6 | 143 | Glycosyltransferase family 10 (fucosyltransferase) C-term. This is the C-terminal domain of a family of fucosyltransferases. This enzyme transfers fucose from GDP-Fucose to GlcNAc in an alpha1,3 linkage. This family is known as glycosyltransferase family 10. The C-terminal domain is the likely binding-region for ADP (manuscript in publication). |
| 215596 | PLN03133 | 2.91e-11 | 83 | 306 | 401 | 607 | beta-1,3-galactosyltransferase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 5.72e-19 | 618 | 766 | 121 | 264 | |
| 2.45e-18 | 62 | 300 | 58 | 284 | |
| 3.35e-18 | 62 | 300 | 83 | 309 | |
| 3.35e-18 | 62 | 300 | 83 | 309 | |
| 7.19e-18 | 613 | 762 | 124 | 267 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.97e-15 | 618 | 765 | 182 | 314 | Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori],2NZW_B Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori],2NZW_C Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori],2NZX_A Crystal Structure of alpha1,3-Fucosyltransferase with GDP [Helicobacter pylori],2NZX_B Crystal Structure of alpha1,3-Fucosyltransferase with GDP [Helicobacter pylori],2NZX_C Crystal Structure of alpha1,3-Fucosyltransferase with GDP [Helicobacter pylori],2NZY_A Crystal Structure of alpha1,3-Fucosyltransferase with GDP-fucose [Helicobacter pylori],2NZY_B Crystal Structure of alpha1,3-Fucosyltransferase with GDP-fucose [Helicobacter pylori],2NZY_C Crystal Structure of alpha1,3-Fucosyltransferase with GDP-fucose [Helicobacter pylori] |
|
| 2.40e-14 | 618 | 765 | 182 | 314 | Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori],5ZOI_B Crystal Structure of alpha1,3-Fucosyltransferase [Helicobacter pylori] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.95e-19 | 62 | 300 | 83 | 309 | Lactosylceramide 1,3-N-acetyl-beta-D-glucosaminyltransferase OS=Xenopus tropicalis OX=8364 GN=b3gnt5 PE=2 SV=1 |
|
| 4.11e-17 | 542 | 762 | 360 | 566 | Putative fucosyltransferase R654 OS=Acanthamoeba polyphaga mimivirus OX=212035 GN=MIMI_R654 PE=3 SV=1 |
|
| 1.54e-16 | 82 | 323 | 93 | 318 | Beta-1,3-galactosyltransferase 1 OS=Gorilla gorilla gorilla OX=9595 GN=B3GALT1 PE=3 SV=1 |
|
| 1.54e-16 | 65 | 300 | 86 | 309 | Lactosylceramide 1,3-N-acetyl-beta-D-glucosaminyltransferase B OS=Xenopus laevis OX=8355 GN=b3gnt5-b PE=2 SV=1 |
|
| 1.54e-16 | 82 | 323 | 93 | 318 | Beta-1,3-galactosyltransferase 1 OS=Pongo pygmaeus OX=9600 GN=B3GALT1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
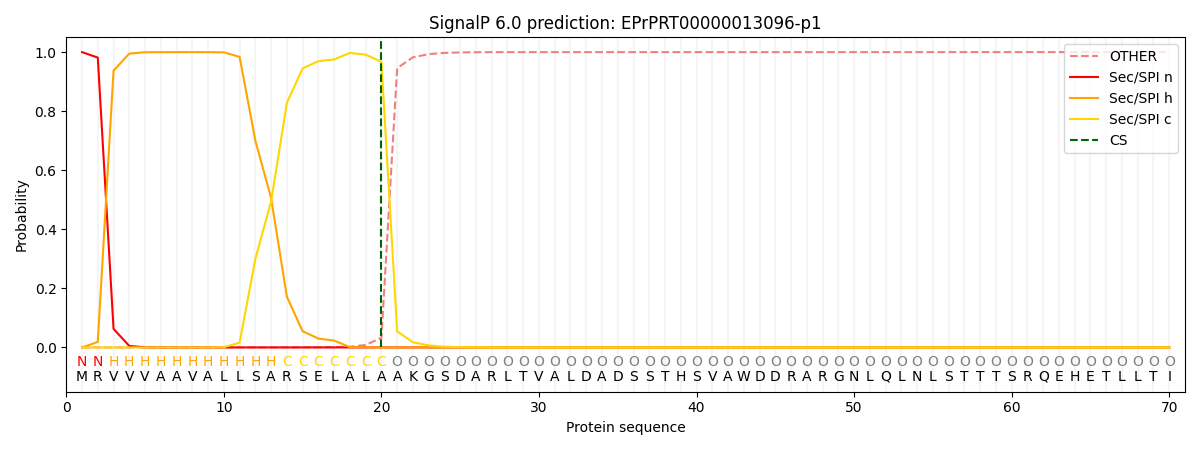
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000328 | 0.999604 | CS pos: 20-21. Pr: 0.9672 |
