You are browsing environment: FUNGIDB
CAZyme Information: EPrPIT00000015584-p1
You are here: Home > Sequence: EPrPIT00000015584-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Globisporangium irregulare | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Pythiaceae; Globisporangium; Globisporangium irregulare | |||||||||||
| CAZyme ID | EPrPIT00000015584-p1 | |||||||||||
| CAZy Family | AA17 | |||||||||||
| CAZyme Description | Callose synthase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.34:28 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT48 | 854 | 1575 | 6.2e-259 | 0.918809201623816 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396784 | Glucan_synthase | 6.05e-174 | 859 | 1539 | 7 | 700 | 1,3-beta-glucan synthase component. This family consists of various 1,3-beta-glucan synthase components including Gls1, Gls2 and Gls3 from yeast. 1,3-beta-glucan synthase EC:2.4.1.34 also known as callose synthase catalyzes the formation of a beta-1,3-glucan polymer that is a major component of the fungal cell wall. The reaction catalyzed is:- UDP-glucose + {(1,3)-beta-D-glucosyl}(N) <=> UDP + {(1,3)-beta-D-glucosyl}(N+1). |
| 405046 | FKS1_dom1 | 1.94e-35 | 150 | 247 | 4 | 106 | 1,3-beta-glucan synthase subunit FKS1, domain-1. The FKS1_dom1 domain is likely to be the 'Class I' region just N-terminal to the first set of transmembrane helices that is involved in 1,3-beta-glucan synthesis itself. This family is found on proteins with family Glucan_synthase, pfam02364. |
| 340916 | MFS_GLUT6_8_Class3_like | 4.90e-14 | 1833 | 2097 | 63 | 322 | Glucose transporter (GLUT) types 6 and 8, Class 3 GLUTs, and similar transporters of the Major Facilitator Superfamily. This subfamily is composed of glucose transporter type 6 (GLUT6), GLUT8, plant early dehydration-induced gene ERD6-like proteins, and similar insect proteins including facilitated trehalose transporter Tret1-1. GLUTs, also called Solute carrier family 2, facilitated glucose transporters (SLC2A), are a family of proteins that facilitate the transport of hexoses such as glucose and fructose. There are fourteen GLUTs found in humans; they display different substrate specificities and tissue expression. They have been categorized into three classes based on sequence similarity: Class 1 (GLUTs 1-4, 14); Class 2 (GLUTs 5, 7, 9, and 11); and Class 3 (GLUTs 6, 8, 10, 12, and HMIT). Insect Tret1-1 is a low-capacity facilitative transporter for trehalose that mediates the transport of trehalose synthesized in the fat body and the incorporation of trehalose into other tissues that require a carbon source. GLUT proteins are comprised of about 500 amino acid residues, possess a single N-linked oligosaccharide, and have 12 transmembrane segments. They belong to the Glucose transporter -like (GLUT-like) family of the Major Facilitator Superfamily (MFS) of membrane transport proteins. MFS proteins are thought to function through a single substrate binding site, alternating-access mechanism involving a rocker-switch type of movement. |
| 395036 | Sugar_tr | 1.11e-13 | 1833 | 2124 | 68 | 362 | Sugar (and other) transporter. |
| 340917 | MFS_XylE_like | 1.41e-08 | 1833 | 1971 | 58 | 194 | D-xylose-proton symporter and similar transporters of the Major Facilitator Superfamily. This subfamily includes bacterial transporters such as D-xylose-proton symporter (XylE or XylT), arabinose-proton symporter (AraE), galactose-proton symporter (GalP), major myo-inositol transporter IolT, glucose transport protein, putative metabolite transport proteins YfiG, YncC, and YwtG, and similar proteins. The symporters XylE, AraE, and GalP facilitate the uptake of D-xylose, arabinose, and galactose, respectively, across the boundary membrane with the concomitant transport of protons into the cell. IolT is involved in polyol metabolism and myo-inositol degradation into acetyl-CoA. The XylE-like subfamily belongs to the Glucose transporter -like (GLUT-like) family of the Major Facilitator Superfamily (MFS) of membrane transport proteins. MFS proteins are thought to function through a single substrate binding site, alternating-access mechanism involving a rocker-switch type of movement. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 29 | 2242 | 21 | 2304 | |
| 0.0 | 25 | 2228 | 14 | 2270 | |
| 0.0 | 68 | 2229 | 43 | 2316 | |
| 0.0 | 30 | 2172 | 12 | 2225 | |
| 0.0 | 70 | 2091 | 69 | 2060 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.31e-06 | 1936 | 2076 | 157 | 304 | The inward-facing structure of the glucose transporter from Staphylococcus epidermidis [Staphylococcus epidermidis ATCC 12228],4LDS_B The inward-facing structure of the glucose transporter from Staphylococcus epidermidis [Staphylococcus epidermidis ATCC 12228] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.78e-230 | 58 | 1752 | 229 | 1910 | Callose synthase 5 OS=Arabidopsis thaliana OX=3702 GN=CALS5 PE=1 SV=1 |
|
| 2.21e-224 | 68 | 1752 | 256 | 1915 | Callose synthase 7 OS=Arabidopsis thaliana OX=3702 GN=CALS7 PE=3 SV=3 |
|
| 9.49e-224 | 68 | 1752 | 233 | 1931 | Callose synthase 1 OS=Arabidopsis thaliana OX=3702 GN=CALS1 PE=1 SV=2 |
|
| 4.60e-222 | 68 | 1752 | 233 | 1931 | Callose synthase 2 OS=Arabidopsis thaliana OX=3702 GN=CALS2 PE=2 SV=3 |
|
| 2.45e-220 | 68 | 1752 | 237 | 1936 | Callose synthase 3 OS=Arabidopsis thaliana OX=3702 GN=CALS3 PE=3 SV=3 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
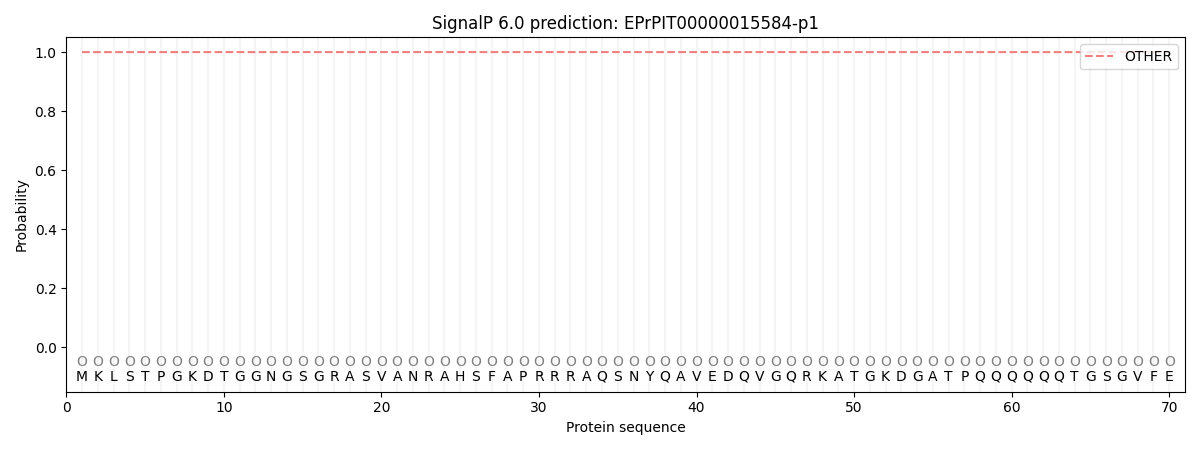
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000052 | 0.000000 |
TMHMM Annotations download full data without filtering help
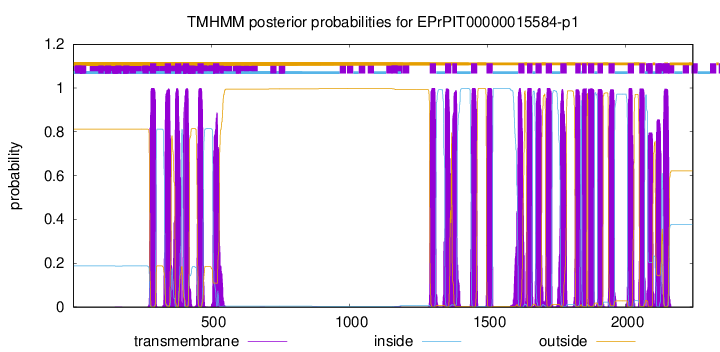
| Start | End |
|---|---|
| 1675 | 1697 |
| 1712 | 1734 |
| 1765 | 1787 |
| 1819 | 1838 |
| 1843 | 1860 |
| 1865 | 1887 |
| 1900 | 1919 |
| 1939 | 1961 |
| 2009 | 2031 |
| 2051 | 2070 |
| 2082 | 2099 |
| 2109 | 2131 |
| 2138 | 2160 |
| 278 | 300 |
| 332 | 354 |
| 369 | 391 |
| 398 | 420 |
| 450 | 472 |
| 507 | 529 |
| 1292 | 1314 |
| 1347 | 1369 |
| 1373 | 1390 |
| 1441 | 1463 |
| 1497 | 1519 |
| 1612 | 1634 |
| 1644 | 1663 |
