You are browsing environment: FUNGIDB
CAZyme Information: EPrPAT00000015190-p1
You are here: Home > Sequence: EPrPAT00000015190-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
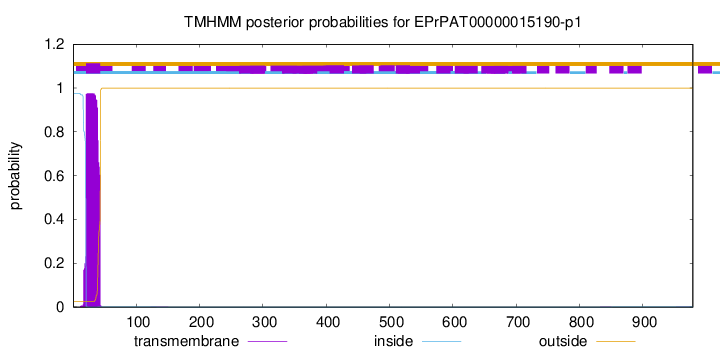
TMHMM annotations
Basic Information help
| Species | Pythium aphanidermatum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Oomycota; NA; ; Pythiaceae; Pythium; Pythium aphanidermatum | |||||||||||
| CAZyme ID | EPrPAT00000015190-p1 | |||||||||||
| CAZy Family | AA3 | |||||||||||
| CAZyme Description | Methyltransferase. | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 27402; End:30777 Strand: - | |||||||||||
Full Sequence Download help
| MVVMMTSEFV RASPWRALLR FALIAATAAV AALAPWNVAV GAVHSRPRVH PIVEIVFPPN | 60 |
| DYVLASSELH IEVAMRIDRL RVPVQGQRVC LAMKTVFAPQ GAPQEAGETV LKETCYEREG | 120 |
| NYTTFHATGL VPGIKYSVSA GLQNAGSMAG YSMRLFEVAS LLLQVDGTTH RMGIPEAMDL | 180 |
| AVDLHVHRDA AKAVEIYRNV LHMLPKHPGA MFRLGQAYLQ DGFPEKAMAL VQDAIQYESS | 240 |
| DPRMYLTMAL CYQAQERHED AILYLERALE LRPSYVFAAL RLGHSRTQVG DWERAIEQYK | 300 |
| AVVATMNEQQ GPQEVLEKPH DSMVVEALAW LCELVRLTNG WYESERCLSQ AVDGFPDRAP | 360 |
| FRIDHGHLLL YSGQFERAMQ EYDAAANLGS IYGKIYAADV LESVGEYEAS IKRYNQARRT | 420 |
| QQAIYDDEHN HDKSLLNLVA VMATTVLPRI LPGSQGAIDD ERERLLTNVE GVIDSIQPDM | 480 |
| NAIPIDPSRI AFSTAVTVNS HNRINKDLKR KIGKMYNMLF IEHRVVNANT APYGVTPMPY | 540 |
| RQYQKRAPPH LSPTHRRLRV GFASRFFYNE AVGLYMNELL PQLNDSKYEI FAFAIGMSKS | 600 |
| LKKYEPVAQI AENVIAVPRE LRIARAEILA ADLDVLIYPE LGMDKTTYFL SYHRLAPIQA | 660 |
| VWWGNPDTSG IPSIDYFITS EHEHESFRNH YTEEVYQMKG MGIYHELPPL PNTTITRQDI | 720 |
| RRAIQERFNV PSDFHYYLCI ESLLHIHPDF DEAIKRLFHR DPHAHIFLLS SSSRKNWKHL | 780 |
| LYDRMLKHIG FEYEHRITFF SDIDTRQEIH LLRAADAVLG SLHMTRPHAS MQAFRAGVPV | 840 |
| VTMPGEMWST RITAGFYKQM DIQELTAGTL DEFVGIAFRL ATDESFQSRM TRKIRRHRGR | 900 |
| LSKDRRAVEE WEKFLDMVAS KTDLALVHDQ PDPESTSTLP HDSTHPSRHP PPTASNDVAL | 960 |
| TCPATTFYHL LSSNFKVLK | 979 |
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT41 | 321 | 930 | 1.7e-64 | 0.573049645390071 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 226428 | Spy | 1.01e-26 | 546 | 898 | 248 | 592 | Predicted O-linked N-acetylglucosamine transferase, SPINDLY family [Posttranslational modification, protein turnover, chaperones]. |
| 276809 | TPR | 9.21e-18 | 210 | 304 | 3 | 97 | Tetratricopeptide repeat. The Tetratricopeptide repeat (TPR) typically contains 34 amino acids and is found in a variety of organisms including bacteria, cyanobacteria, yeast, fungi, plants, and humans. It is present in a variety of proteins including those involved in chaperone, cell-cycle, transcription, and protein transport complexes. The number of TPR motifs varies among proteins. Those containing 5-6 tandem repeats generate a right-handed helical structure with an amphipathic channel that is thought to accommodate an alpha-helix of a target protein. It has been proposed that TPR proteins preferentially interact with WD-40 repeat proteins, but in many instances several TPR-proteins seem to aggregate to multi-protein complexes. |
| 276809 | TPR | 1.75e-14 | 180 | 270 | 7 | 97 | Tetratricopeptide repeat. The Tetratricopeptide repeat (TPR) typically contains 34 amino acids and is found in a variety of organisms including bacteria, cyanobacteria, yeast, fungi, plants, and humans. It is present in a variety of proteins including those involved in chaperone, cell-cycle, transcription, and protein transport complexes. The number of TPR motifs varies among proteins. Those containing 5-6 tandem repeats generate a right-handed helical structure with an amphipathic channel that is thought to accommodate an alpha-helix of a target protein. It has been proposed that TPR proteins preferentially interact with WD-40 repeat proteins, but in many instances several TPR-proteins seem to aggregate to multi-protein complexes. |
| 223533 | TPR | 9.32e-10 | 189 | 300 | 145 | 260 | Tetratricopeptide (TPR) repeat [General function prediction only]. |
| 276809 | TPR | 1.86e-09 | 242 | 303 | 1 | 62 | Tetratricopeptide repeat. The Tetratricopeptide repeat (TPR) typically contains 34 amino acids and is found in a variety of organisms including bacteria, cyanobacteria, yeast, fungi, plants, and humans. It is present in a variety of proteins including those involved in chaperone, cell-cycle, transcription, and protein transport complexes. The number of TPR motifs varies among proteins. Those containing 5-6 tandem repeats generate a right-handed helical structure with an amphipathic channel that is thought to accommodate an alpha-helix of a target protein. It has been proposed that TPR proteins preferentially interact with WD-40 repeat proteins, but in many instances several TPR-proteins seem to aggregate to multi-protein complexes. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| UIZ20641.1|GT41 | 3.20e-166 | 51 | 916 | 36 | 901 |
| QTA91146.1|GT41 | 3.02e-57 | 558 | 921 | 509 | 866 |
| BAZ61948.1|GT41 | 6.47e-54 | 447 | 923 | 448 | 889 |
| BAZ15802.1|GT41 | 6.47e-54 | 447 | 923 | 448 | 889 |
| BBO88349.1|GT41 | 1.93e-51 | 557 | 921 | 522 | 875 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5A01_A | 4.06e-09 | 553 | 894 | 230 | 668 | O-GlcNAc transferase from Drososphila melanogaster [Drosophila melanogaster],5A01_B O-GlcNAc transferase from Drososphila melanogaster [Drosophila melanogaster],5A01_C O-GlcNAc transferase from Drososphila melanogaster [Drosophila melanogaster] |
| 3Q3E_A | 3.04e-07 | 639 | 899 | 356 | 604 | Crystal structure of the Actinobacillus pleuropneumoniae HMW1C glycosyltransferase [Actinobacillus pleuropneumoniae serovar 1 str. 4074],3Q3E_B Crystal structure of the Actinobacillus pleuropneumoniae HMW1C glycosyltransferase [Actinobacillus pleuropneumoniae serovar 1 str. 4074],3Q3H_A Crystal structure of the Actinobacillus pleuropneumoniae HMW1C glycosyltransferase in complex with UDP-GLC [Actinobacillus pleuropneumoniae serovar 1 str. 4074],3Q3H_B Crystal structure of the Actinobacillus pleuropneumoniae HMW1C glycosyltransferase in complex with UDP-GLC [Actinobacillus pleuropneumoniae serovar 1 str. 4074],3Q3I_A Crystal structure of the Actinobacillus pleuropneumoniae HMW1C glycosyltransferase in the presence of peptide N1131 [Actinobacillus pleuropneumoniae serovar 1 str. 4074],3Q3I_B Crystal structure of the Actinobacillus pleuropneumoniae HMW1C glycosyltransferase in the presence of peptide N1131 [Actinobacillus pleuropneumoniae serovar 1 str. 4074] |
| 2JLB_A | 1.49e-06 | 551 | 883 | 195 | 522 | Xanthomonas campestris putative OGT (XCC0866), complex with UDP- GlcNAc phosphonate analogue [Xanthomonas campestris pv. campestris],2JLB_B Xanthomonas campestris putative OGT (XCC0866), complex with UDP- GlcNAc phosphonate analogue [Xanthomonas campestris pv. campestris],2VSY_A Xanthomonas campestris putative OGT (XCC0866), apostructure [Xanthomonas campestris pv. campestris str. ATCC 33913],2VSY_B Xanthomonas campestris putative OGT (XCC0866), apostructure [Xanthomonas campestris pv. campestris str. ATCC 33913],2XGM_A Substrate and product analogues as human O-GlcNAc transferase inhibitors. [Xanthomonas campestris],2XGM_B Substrate and product analogues as human O-GlcNAc transferase inhibitors. [Xanthomonas campestris],2XGO_A XcOGT in complex with UDP-S-GlcNAc [Xanthomonas campestris],2XGO_B XcOGT in complex with UDP-S-GlcNAc [Xanthomonas campestris],2XGS_A XcOGT in complex with C-UDP [Xanthomonas campestris],2XGS_B XcOGT in complex with C-UDP [Xanthomonas campestris] |
| 2VSN_A | 1.49e-06 | 551 | 883 | 195 | 522 | Structure and topological arrangement of an O-GlcNAc transferase homolog: insight into molecular control of intracellular glycosylation [Xanthomonas campestris pv. campestris str. 8004],2VSN_B Structure and topological arrangement of an O-GlcNAc transferase homolog: insight into molecular control of intracellular glycosylation [Xanthomonas campestris pv. campestris str. 8004] |
| 6Q4M_A | 8.60e-06 | 548 | 898 | 228 | 683 | Crystal structure of the O-GlcNAc transferase Asn648Tyr mutation [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| sp|Q9M8Y0|SEC_ARATH | 3.07e-10 | 556 | 898 | 590 | 929 | Probable UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase SEC OS=Arabidopsis thaliana OX=3702 GN=SEC PE=1 SV=1 |
| sp|A3N2T3|NGT_ACTP2 | 1.55e-06 | 639 | 899 | 345 | 593 | UDP-glucose:protein N-beta-glucosyltransferase OS=Actinobacillus pleuropneumoniae serotype 5b (strain L20) OX=416269 GN=APL_1635 PE=1 SV=1 |
| sp|O82039|SPY_PETHY | 4.15e-06 | 654 | 895 | 588 | 820 | Probable UDP-N-acetylglucosamine--peptide N-acetylglucosaminyltransferase SPINDLY OS=Petunia hybrida OX=4102 GN=SPY PE=2 SV=1 |
| sp|B3H2N2|NGT_ACTP7 | 6.10e-06 | 639 | 899 | 345 | 593 | UDP-glucose:protein N-beta-glucosyltransferase OS=Actinobacillus pleuropneumoniae serotype 7 (strain AP76) OX=537457 GN=APP7_1697 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
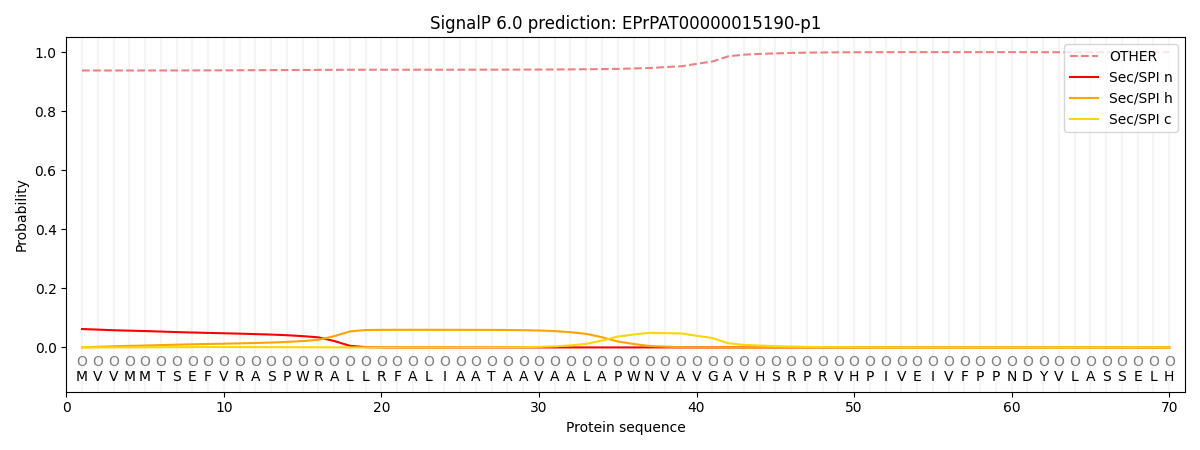
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.941323 | 0.058707 |