You are browsing environment: FUNGIDB
CAZyme Information: ELR04211.1
You are here: Home > Sequence: ELR04211.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Pseudogymnoascus destructans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Leotiomycetes; ; Pseudeurotiaceae; Pseudogymnoascus; Pseudogymnoascus destructans | |||||||||||
| CAZyme ID | ELR04211.1 | |||||||||||
| CAZy Family | GH13 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT15 | 43 | 325 | 2.1e-109 | 0.9010989010989011 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 227353 | KTR1 | 3.17e-118 | 43 | 371 | 93 | 392 | Mannosyltransferase [Carbohydrate transport and metabolism]. |
| 396385 | Glyco_transf_15 | 4.63e-95 | 42 | 325 | 54 | 300 | Glycolipid 2-alpha-mannosyltransferase. This is a family of alpha-1,2 mannosyl-transferases involved in N-linked and O-linked glycosylation of proteins. Some of the enzymes in this family have been shown to be involved in O- and N-linked glycan modifications in the Golgi. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.10e-216 | 1 | 375 | 60 | 504 | |
| 1.17e-215 | 1 | 375 | 60 | 504 | |
| 1.17e-215 | 1 | 375 | 60 | 504 | |
| 1.66e-215 | 1 | 375 | 60 | 504 | |
| 4.14e-210 | 1 | 375 | 62 | 499 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.41e-69 | 43 | 370 | 39 | 336 | Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p [Saccharomyces cerevisiae],1S4N_B Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p [Saccharomyces cerevisiae],1S4O_A Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p: binary complex with GDP/Mn [Saccharomyces cerevisiae],1S4O_B Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p: binary complex with GDP/Mn [Saccharomyces cerevisiae],1S4P_A Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p: ternary complex with GDP/Mn and methyl-alpha-mannoside acceptor [Saccharomyces cerevisiae],1S4P_B Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p: ternary complex with GDP/Mn and methyl-alpha-mannoside acceptor [Saccharomyces cerevisiae] |
|
| 2.15e-57 | 43 | 331 | 79 | 338 | Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_B Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_C Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_D Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_E Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_F Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOP_A Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_B Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_C Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_D Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_E Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_F Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163] |
|
| 5.35e-49 | 43 | 366 | 115 | 422 | X-ray structure of the mannosyltransferase Ktr4p from S. cerevisiae in complex with GDP [Saccharomyces cerevisiae S288C],5A07_B X-ray structure of the mannosyltransferase Ktr4p from S. cerevisiae in complex with GDP [Saccharomyces cerevisiae S288C],5A08_A X-ray structure of the mannosyltransferase Ktr4p from S. cerevisiae [Saccharomyces cerevisiae S288C],5A08_B X-ray structure of the mannosyltransferase Ktr4p from S. cerevisiae [Saccharomyces cerevisiae S288C] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.04e-109 | 43 | 370 | 104 | 434 | O-glycoside alpha-1,2-mannosyltransferase homolog 4 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=omh4 PE=3 SV=2 |
|
| 3.18e-98 | 43 | 372 | 96 | 418 | O-glycoside alpha-1,2-mannosyltransferase homolog 5 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=omh5 PE=3 SV=2 |
|
| 6.61e-85 | 43 | 371 | 100 | 484 | Probable mannosyltransferase KTR7 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=KTR7 PE=1 SV=1 |
|
| 5.89e-84 | 43 | 321 | 97 | 407 | Probable mannosyltransferase KTR5 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=KTR5 PE=1 SV=1 |
|
| 9.28e-68 | 43 | 370 | 133 | 430 | Glycolipid 2-alpha-mannosyltransferase OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=KRE2 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
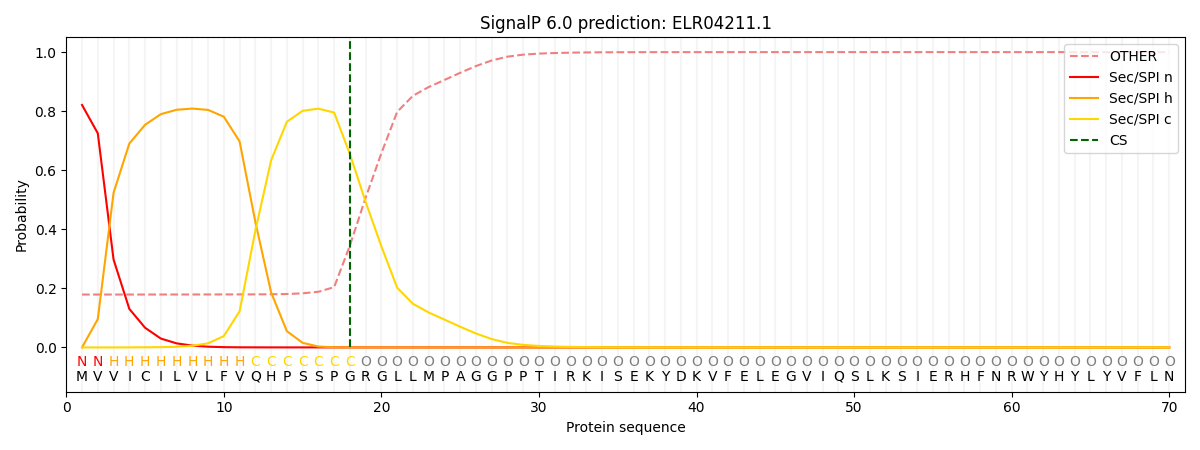
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.185353 | 0.814619 | CS pos: 18-19. Pr: 0.6547 |
