You are browsing environment: FUNGIDB
CAZyme Information: EKG19809.1
You are here: Home > Sequence: EKG19809.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Macrophomina phaseolina | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Dothideomycetes; ; Botryosphaeriaceae; Macrophomina; Macrophomina phaseolina | |||||||||||
| CAZyme ID | EKG19809.1 | |||||||||||
| CAZy Family | GH92 | |||||||||||
| CAZyme Description | Glycoside hydrolase family 10 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.8:45 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH10 | 29 | 322 | 1.9e-106 | 0.9933993399339934 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395262 | Glyco_hydro_10 | 2.12e-144 | 29 | 321 | 1 | 310 | Glycosyl hydrolase family 10. |
| 214750 | Glyco_10 | 4.72e-119 | 72 | 319 | 1 | 263 | Glycosyl hydrolase family 10. |
| 226217 | XynA | 3.60e-74 | 7 | 320 | 3 | 338 | Endo-1,4-beta-xylanase, GH35 family [Carbohydrate transport and metabolism]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.36e-165 | 23 | 323 | 22 | 323 | |
| 3.36e-165 | 23 | 323 | 22 | 323 | |
| 3.36e-165 | 23 | 323 | 22 | 323 | |
| 3.36e-165 | 23 | 323 | 22 | 323 | |
| 1.36e-164 | 23 | 323 | 22 | 323 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.95e-149 | 25 | 323 | 2 | 302 | Crystal structure of an endo-beta-1,4-xylanase (glycoside hydrolase family 10/GH10) enzyme from Aspergillus niger [Aspergillus niger CBS 513.88],4XUY_B Crystal structure of an endo-beta-1,4-xylanase (glycoside hydrolase family 10/GH10) enzyme from Aspergillus niger [Aspergillus niger CBS 513.88] |
|
| 1.62e-147 | 25 | 323 | 2 | 302 | XYLANASE FROM PENICILLIUM SIMPLICISSIMUM, COMPLEX WITH 1,2-(4-DEOXY-BETA-L-THREO-HEX-4-ENOPYRANOSYLURONIC ACID)-BETA-1,4-XYLOTRIOSE) [Penicillium simplicissimum],1B31_A XYLANASE FROM PENICILLIUM SIMPLICISSIMUM, NATIVE WITH PEG200 AS CRYOPROTECTANT [Penicillium simplicissimum],1B3V_A XYLANASE FROM PENICILLIUM SIMPLICISSIMUM, COMPLEX WITH XYLOSE [Penicillium simplicissimum],1B3W_A XYLANASE FROM PENICILLIUM SIMPLICISSIMUM, COMPLEX WITH XYLOBIOSE [Penicillium simplicissimum],1B3X_A XYLANASE FROM PENICILLIUM SIMPLICISSIMUM, COMPLEX WITH XYLOTRIOSE [Penicillium simplicissimum],1B3Y_A XYLANASE FROM PENICILLIUM SIMPLICISSIMUM, COMPLEX WITH XYLOTETRAOSE [Penicillium simplicissimum],1B3Z_A XYLANASE FROM PENICILLIUM SIMPLICISSIMUM, COMPLEX WITH XYLOPENTAOSE [Penicillium simplicissimum],1BG4_A XYLANASE FROM PENICILLIUM SIMPLICISSIMUM [Penicillium simplicissimum] |
|
| 4.81e-147 | 24 | 323 | 1 | 301 | Structural studies on the mobility in the active site of the Thermoascus aurantiacus xylanase I [Thermoascus aurantiacus] |
|
| 2.77e-146 | 25 | 323 | 2 | 301 | Thermostable xylanase I from Thermoascus aurantiacus- Crystal form II [Thermoascus aurantiacus],1GOM_A Thermostable xylanase I from Thermoascus aurantiacus- Crystal form I [Thermoascus aurantiacus],1GOO_A Thermostable xylanase I from Thermoascus aurantiacus - Cryocooled glycerol complex [Thermoascus aurantiacus],1GOQ_A Thermostable xylanase I from Thermoascus aurantiacus - Room temperature xylobiose complex [Thermoascus aurantiacus],1GOR_A THERMOSTABLE XYLANASE I FROM THERMOASCUS AURANTIACUS - XYLOBIOSE COMPLEX AT 100 K [Thermoascus aurantiacus] |
|
| 9.18e-145 | 28 | 323 | 4 | 302 | Crystal Structure of xylanase (GH10) in complex with inhibitor (XIP) [Aspergillus nidulans] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.96e-166 | 23 | 323 | 22 | 323 | Endo-1,4-beta-xylanase F3 OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=xynF3 PE=1 SV=1 |
|
| 2.37e-152 | 16 | 323 | 17 | 327 | Endo-1,4-beta-xylanase F1 OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=xynF1 PE=1 SV=1 |
|
| 3.91e-151 | 6 | 323 | 8 | 327 | Probable endo-1,4-beta-xylanase C OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=xlnC PE=1 SV=1 |
|
| 5.35e-151 | 23 | 323 | 25 | 326 | Probable endo-1,4-beta-xylanase C OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=xlnC PE=2 SV=2 |
|
| 1.12e-150 | 4 | 323 | 6 | 327 | Endo-1,4-beta-xylanase A OS=Penicillium canescens OX=5083 GN=xylA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
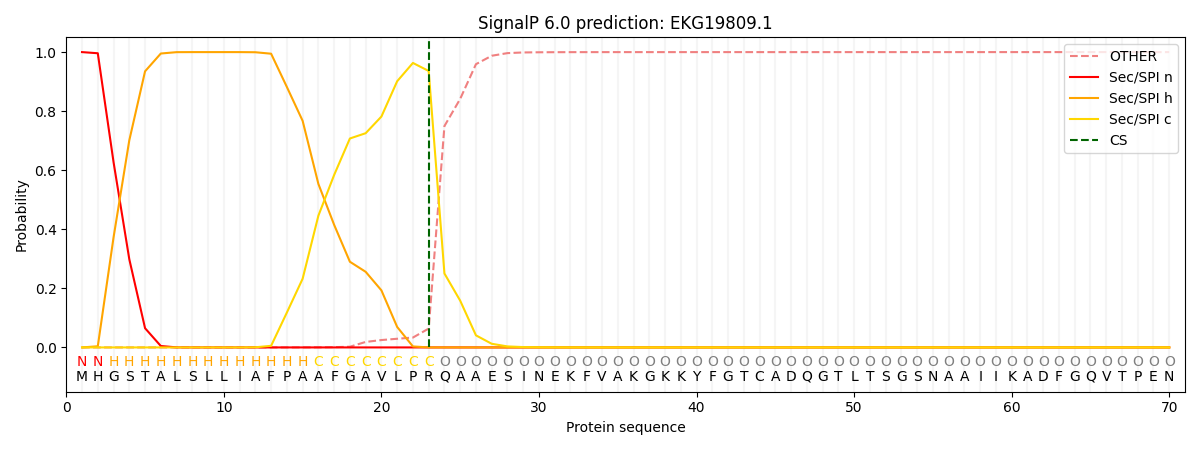
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000214 | 0.999767 | CS pos: 23-24. Pr: 0.9361 |
