You are browsing environment: FUNGIDB
CAZyme Information: EJT78090.1
You are here: Home > Sequence: EJT78090.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Gaeumannomyces tritici | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Magnaporthaceae; Gaeumannomyces; Gaeumannomyces tritici | |||||||||||
| CAZyme ID | EJT78090.1 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | Beta-glucosidase [Source:UniProtKB/TrEMBL;Acc:J3NPI5] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.21:13 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 92 | 306 | 1.5e-61 | 0.9861111111111112 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 224389 | BglX | 2.09e-53 | 93 | 440 | 55 | 370 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| 396478 | Glyco_hydro_3_C | 1.24e-50 | 409 | 639 | 1 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
| 185053 | PRK15098 | 9.91e-39 | 115 | 861 | 116 | 756 | beta-glucosidase BglX. |
| 395747 | Glyco_hydro_3 | 2.56e-38 | 117 | 346 | 86 | 313 | Glycosyl hydrolase family 3 N terminal domain. |
| 405066 | Fn3-like | 5.46e-25 | 785 | 859 | 2 | 71 | Fibronectin type III-like domain. This domain has a fibronectin type III-like structure. It is often found in association with pfam00933 and pfam01915. Its function is unknown. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 880 | 1 | 875 | |
| 0.0 | 9 | 881 | 6 | 871 | |
| 0.0 | 9 | 881 | 6 | 871 | |
| 0.0 | 9 | 881 | 6 | 871 | |
| 0.0 | 17 | 881 | 9 | 869 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.38e-217 | 16 | 862 | 7 | 845 | Chain A, Beta-glucosidase [Rasamsonia emersonii],5JU6_B Chain B, Beta-glucosidase [Rasamsonia emersonii],5JU6_C Chain C, Beta-glucosidase [Rasamsonia emersonii],5JU6_D Chain D, Beta-glucosidase [Rasamsonia emersonii] |
|
| 7.67e-217 | 37 | 862 | 5 | 831 | Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus [Aspergillus aculeatus],4IIB_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus [Aspergillus aculeatus],4IIC_A Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with isofagomine [Aspergillus aculeatus],4IIC_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with isofagomine [Aspergillus aculeatus],4IID_A Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with 1-deoxynojirimycin [Aspergillus aculeatus],4IID_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with 1-deoxynojirimycin [Aspergillus aculeatus],4IIE_A Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with calystegine B(2) [Aspergillus aculeatus],4IIE_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with calystegine B(2) [Aspergillus aculeatus],4IIF_A Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with castanospermine [Aspergillus aculeatus],4IIF_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with castanospermine [Aspergillus aculeatus],4IIG_A Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with D-glucose [Aspergillus aculeatus],4IIG_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with D-glucose [Aspergillus aculeatus],4IIH_A Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with thiocellobiose [Aspergillus aculeatus],4IIH_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with thiocellobiose [Aspergillus aculeatus] |
|
| 2.51e-215 | 37 | 862 | 6 | 832 | Three-dimensional structures of two heavily N-glycosylated Aspergillus sp. Family GH3 beta-D-glucosidases [Aspergillus oryzae],5FJJ_B Three-dimensional structures of two heavily N-glycosylated Aspergillus sp. Family GH3 beta-D-glucosidases [Aspergillus oryzae],5FJJ_C Three-dimensional structures of two heavily N-glycosylated Aspergillus sp. Family GH3 beta-D-glucosidases [Aspergillus oryzae],5FJJ_D Three-dimensional structures of two heavily N-glycosylated Aspergillus sp. Family GH3 beta-D-glucosidases [Aspergillus oryzae] |
|
| 2.63e-210 | 38 | 862 | 4 | 829 | Chain A, Beta-glucosidase [Thermochaetoides thermophila] |
|
| 6.75e-207 | 37 | 862 | 5 | 834 | Three-dimensional structures of two heavily N-glycosylated Aspergillus sp. Family GH3 beta-D-glucosidases [Aspergillus fumigatus],5FJI_B Three-dimensional structures of two heavily N-glycosylated Aspergillus sp. Family GH3 beta-D-glucosidases [Aspergillus fumigatus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 8 | 866 | 7 | 857 | Probable beta-glucosidase F OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=bglF PE=3 SV=1 |
|
| 0.0 | 9 | 866 | 10 | 860 | Probable beta-glucosidase F OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=bglF PE=3 SV=2 |
|
| 0.0 | 8 | 866 | 7 | 857 | Probable beta-glucosidase F OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=bglF PE=3 SV=1 |
|
| 0.0 | 9 | 866 | 8 | 859 | Probable beta-glucosidase F OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=bglF PE=3 SV=2 |
|
| 0.0 | 17 | 866 | 18 | 858 | Probable beta-glucosidase F OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=bglF PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
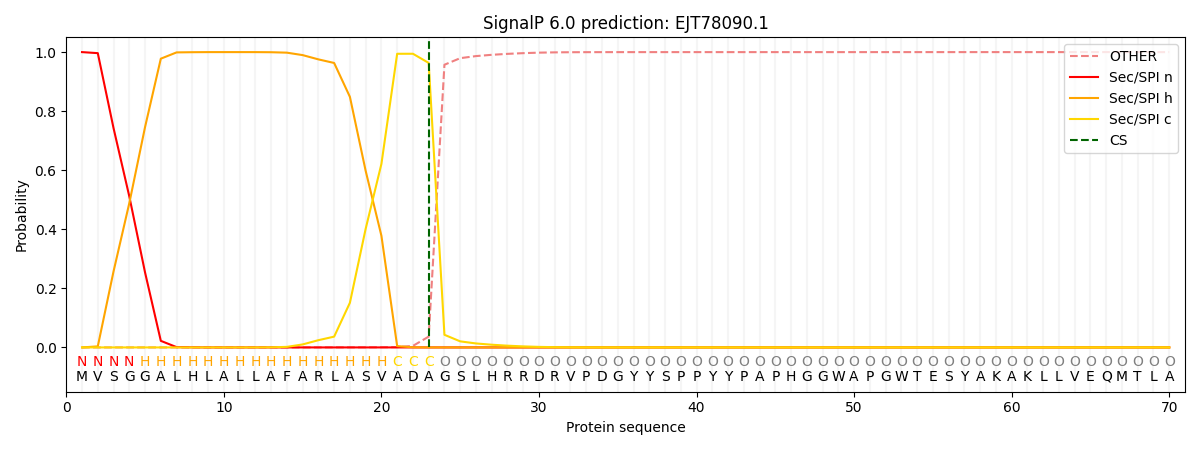
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000286 | 0.999681 | CS pos: 23-24. Pr: 0.9633 |
