You are browsing environment: FUNGIDB
CAZyme Information: EJT73808.1
You are here: Home > Sequence: EJT73808.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
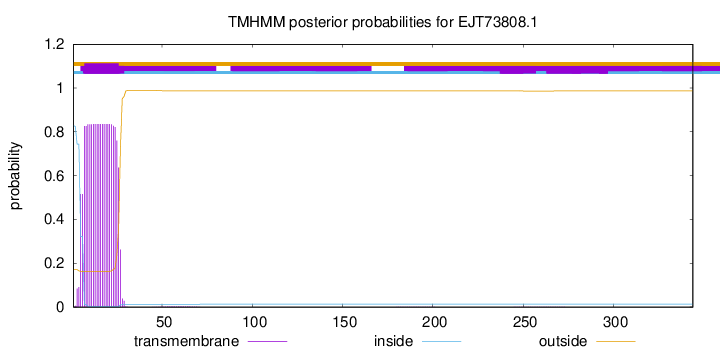
TMHMM annotations
Basic Information help
| Species | Gaeumannomyces tritici | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Magnaporthaceae; Gaeumannomyces; Gaeumannomyces tritici | |||||||||||
| CAZyme ID | EJT73808.1 | |||||||||||
| CAZy Family | GH13 | |||||||||||
| CAZyme Description | CBM1 domain-containing protein [Source:UniProtKB/TrEMBL;Acc:J3P2B6] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH131 | 24 | 264 | 1.9e-109 | 0.996078431372549 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 408085 | GH131_N | 6.14e-79 | 24 | 253 | 1 | 246 | Glycoside hydrolase 131 catalytic N-terminal domain. This is the N-terminal domain found in glycoside hydrolase family 131 (GH131A) protein observed in Coprinopsis cinerea. GH131A exhibits bifunctional exo-beta-1,3-/-1,6- and endo-beta-1,4 activity toward beta-glucan. This domain is catalytic in nature though the catalytic mechanism of C. cinerea GH131A is different from that of typical glycosidases that use a pair of carboxylic acid residues as the catalytic residues. In the case of GH131A, Glu98 and His218 may form a catalytic dyad and Glu98 may activate His218 during catalysis. |
| 197593 | fCBD | 4.93e-10 | 312 | 344 | 1 | 34 | Fungal-type cellulose-binding domain. Small four-cysteine cellulose-binding domain of fungi |
| 395595 | CBM_1 | 2.72e-09 | 313 | 340 | 1 | 29 | Fungal cellulose binding domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.82e-189 | 15 | 343 | 12 | 346 | |
| 3.27e-159 | 7 | 344 | 4 | 363 | |
| 4.65e-159 | 7 | 344 | 4 | 363 | |
| 4.65e-159 | 7 | 344 | 4 | 363 | |
| 4.65e-159 | 7 | 344 | 4 | 363 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.23e-157 | 22 | 261 | 1 | 240 | Chain A, Beta-glucanase [Podospora anserina],4LE3_B Chain B, Beta-glucanase [Podospora anserina],4LE3_C Chain C, Beta-glucanase [Podospora anserina],4LE3_D Chain D, Beta-glucanase [Podospora anserina],4LE4_A Chain A, Beta-glucanase [Podospora anserina],4LE4_B Chain B, Beta-glucanase [Podospora anserina],4LE4_C Chain C, Beta-glucanase [Podospora anserina],4LE4_D Chain D, Beta-glucanase [Podospora anserina] |
|
| 5.31e-81 | 22 | 264 | 1 | 234 | Chain A, Crystal structure of the catalytic domain of the glycoside hydrolase family 131 protein from Coprinopsis cinerea [Coprinopsis cinerea okayama7#130],3W9A_B Chain B, Crystal structure of the catalytic domain of the glycoside hydrolase family 131 protein from Coprinopsis cinerea [Coprinopsis cinerea okayama7#130],3W9A_C Chain C, Crystal structure of the catalytic domain of the glycoside hydrolase family 131 protein from Coprinopsis cinerea [Coprinopsis cinerea okayama7#130],3W9A_D Chain D, Crystal structure of the catalytic domain of the glycoside hydrolase family 131 protein from Coprinopsis cinerea [Coprinopsis cinerea okayama7#130] |
|
| 6.79e-07 | 311 | 344 | 2 | 36 | Chain A, C-TERMINAL DOMAIN OF CELLOBIOHYDROLASE I [Trichoderma reesei],2CBH_A Chain A, C-TERMINAL DOMAIN OF CELLOBIOHYDROLASE I [Trichoderma reesei],2MWJ_A Chain A, Exoglucanase 1 [Trichoderma reesei],2MWK_A Chain A, Exoglucanase 1 [Trichoderma reesei],5X34_A Chain A, Exoglucanase 1 [Trichoderma reesei],5X35_A Chain A, Exoglucanase 1 [Trichoderma reesei],5X36_A Chain A, Exoglucanase 1 [Trichoderma reesei],5X37_A Chain A, Exoglucanase 1 [Trichoderma reesei],5X38_A Chain A, Exoglucanase 1 [Trichoderma reesei] |
|
| 3.26e-06 | 311 | 344 | 2 | 36 | Chain A, CELLOBIOHYDROLASE I [Trichoderma reesei],1AZH_A Chain A, CELLOBIOHYDROLASE I [Trichoderma reesei],5X3C_A Chain A, Exoglucanase 1 [Trichoderma reesei] |
|
| 3.26e-06 | 311 | 344 | 2 | 36 | Chain A, CELLOBIOHYDROLASE I [Trichoderma reesei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.68e-08 | 306 | 344 | 303 | 342 | Polysaccharide monooxygenase Cel61a OS=Myceliophthora thermophila (strain ATCC 42464 / BCRC 31852 / DSM 1799) OX=573729 GN=Cel61a PE=1 SV=1 |
|
| 6.61e-06 | 306 | 344 | 390 | 429 | Endo-1,4-beta-xylanase 1 OS=Humicola grisea var. thermoidea OX=5528 GN=xyn1 PE=2 SV=1 |
|
| 6.87e-06 | 308 | 344 | 18 | 55 | 1,4-beta-D-glucan cellobiohydrolase C OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=cbhC PE=1 SV=2 |
|
| 7.09e-06 | 307 | 344 | 21 | 59 | 4-O-methyl-glucuronoyl methylesterase OS=Podospora anserina (strain S / ATCC MYA-4624 / DSM 980 / FGSC 10383) OX=515849 GN=ge1 PE=1 SV=1 |
|
| 9.76e-06 | 309 | 344 | 478 | 514 | Exoglucanase 1 OS=Hypocrea rufa OX=5547 GN=cbh1 PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
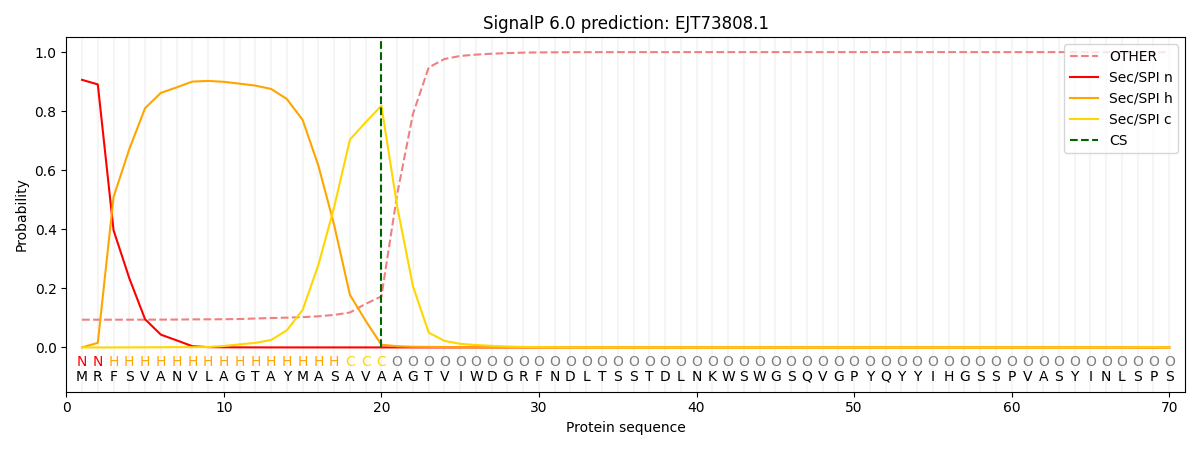
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.103652 | 0.896328 | CS pos: 20-21. Pr: 0.8183 |