You are browsing environment: FUNGIDB
CAZyme Information: EGD93360.1
You are here: Home > Sequence: EGD93360.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Trichophyton tonsurans | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Arthrodermataceae; Trichophyton; Trichophyton tonsurans | |||||||||||
| CAZyme ID | EGD93360.1 | |||||||||||
| CAZy Family | AA5 | |||||||||||
| CAZyme Description | mannosyl-oligosaccharide glucosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.106:25 | 3.2.1.-:4 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH63 | 580 | 791 | 2.3e-30 | 0.43157894736842106 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397353 | Glyco_hydro_63 | 0.0 | 302 | 793 | 1 | 494 | Glycosyl hydrolase family 63 C-terminal domain. This is a family of eukaryotic enzymes belonging to glycosyl hydrolase family 63. They catalyze the specific cleavage of the non-reducing terminal glucose residue from Glc(3)Man(9)GlcNAc(2). Mannosyl oligosaccharide glucosidase EC:3.2.1.106 is the first enzyme in the N-linked oligosaccharide processing pathway. This family represents the C-terminal catalytic domain. |
| 407154 | Glyco_hydro_63N | 9.24e-120 | 38 | 260 | 1 | 221 | Glycosyl hydrolase family 63 N-terminal domain. This is a family of eukaryotic enzymes belonging to glycosyl hydrolase family 63. They catalyze the specific cleavage of the non-reducing terminal glucose residue from Glc(3)Man(9)GlcNAc(2). Mannosyl oligosaccharide glucosidase EC:3.2.1.106 is the first enzyme in the N-linked oligosaccharide processing pathway. This family represents the N-terminal beta sandwich domain. |
| 225942 | GDB1 | 5.83e-07 | 594 | 745 | 428 | 565 | Glycogen debranching enzyme (alpha-1,6-glucosidase) [Carbohydrate transport and metabolism]. |
| 215307 | PLN02567 | 0.006 | 583 | 788 | 328 | 537 | alpha,alpha-trehalase |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 33 | 806 | 43 | 818 | |
| 0.0 | 21 | 806 | 23 | 817 | |
| 0.0 | 19 | 805 | 19 | 816 | |
| 0.0 | 21 | 806 | 23 | 817 | |
| 0.0 | 19 | 805 | 19 | 816 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.87e-177 | 35 | 791 | 12 | 800 | Crystal structure of Processing alpha-Glucosidase I [Saccharomyces cerevisiae S288C] |
|
| 6.75e-139 | 24 | 791 | 20 | 775 | Murine endoplasmic reticulum alpha-glucosidase I with N-9'-methoxynonyl-1-deoxynojirimycin [Mus musculus],5MHF_B Murine endoplasmic reticulum alpha-glucosidase I with N-9'-methoxynonyl-1-deoxynojirimycin [Mus musculus],5MHF_C Murine endoplasmic reticulum alpha-glucosidase I with N-9'-methoxynonyl-1-deoxynojirimycin [Mus musculus],5MHF_D Murine endoplasmic reticulum alpha-glucosidase I with N-9'-methoxynonyl-1-deoxynojirimycin [Mus musculus] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.03e-240 | 30 | 792 | 34 | 805 | Probable mannosyl-oligosaccharide glucosidase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPAC6G10.09 PE=3 SV=1 |
|
| 8.79e-176 | 35 | 791 | 42 | 830 | Mannosyl-oligosaccharide glucosidase OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=CWH41 PE=1 SV=1 |
|
| 4.29e-140 | 24 | 791 | 75 | 830 | Mannosyl-oligosaccharide glucosidase OS=Mus musculus OX=10090 GN=Mogs PE=1 SV=1 |
|
| 8.20e-136 | 24 | 791 | 75 | 830 | Mannosyl-oligosaccharide glucosidase OS=Rattus norvegicus OX=10116 GN=Mogs PE=1 SV=1 |
|
| 2.18e-132 | 17 | 791 | 79 | 833 | Mannosyl-oligosaccharide glucosidase OS=Homo sapiens OX=9606 GN=MOGS PE=1 SV=5 |
SignalP and Lipop Annotations help
This protein is predicted as SP
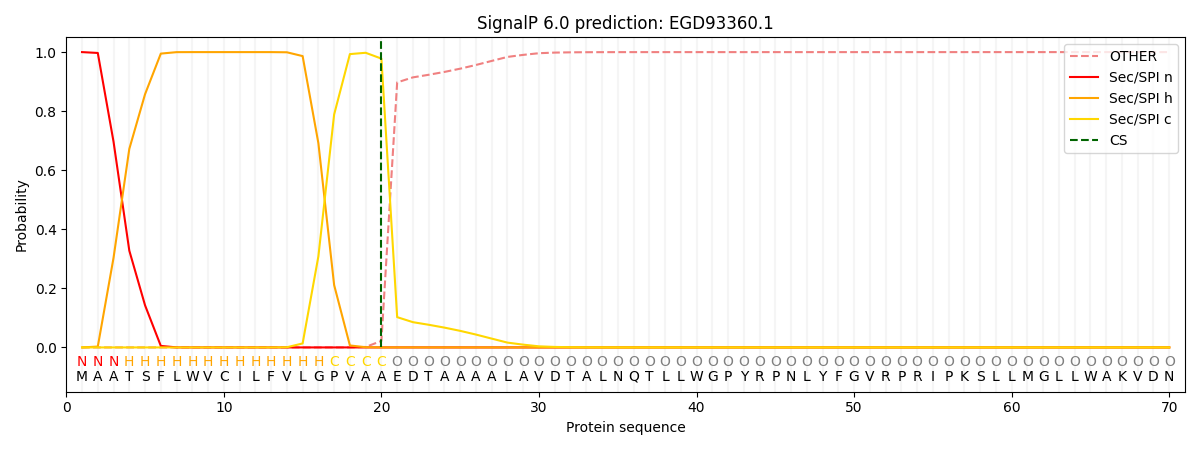
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000243 | 0.999729 | CS pos: 20-21. Pr: 0.9780 |
