You are browsing environment: FUNGIDB
CAZyme Information: EEU47828.1
You are here: Home > Sequence: EEU47828.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium vanettenii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium vanettenii | |||||||||||
| CAZyme ID | EEU47828.1 | |||||||||||
| CAZy Family | GT57 | |||||||||||
| CAZyme Description | glycosyltransferase family 39 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.109:8 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT39 | 56 | 301 | 4.1e-71 | 0.9955156950672646 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 224839 | PMT1 | 0.0 | 49 | 772 | 22 | 699 | Dolichyl-phosphate-mannose--protein O-mannosyl transferase [Posttranslational modification, protein turnover, chaperones]. |
| 396786 | PMT | 1.36e-81 | 55 | 305 | 1 | 245 | Dolichyl-phosphate-mannose-protein mannosyltransferase. This is a family of Dolichyl-phosphate-mannose-protein mannosyltransferase proteins EC:2.4.1.109. These proteins are responsible for O-linked glycosylation of proteins, they catalyze the reaction:- Dolichyl phosphate D-mannose + protein <=> dolichyl phosphate + O-D-mannosyl-protein. Also in this family is Drosophila rotated abdomen protein which is a putative mannosyltransferase. This family appears to be distantly related to pfam02516 (A Bateman pers. obs.). This family also contains sequences from ArnTs (4-amino-4-deoxy-L-arabinose lipid A transferase). They catalyze the addition of 4-amino-4-deoxy-l-arabinose (l-Ara4N) to the lipid A moiety of the lipopolysaccharide. This is a critical modification enabling bacteria (e.g. Escherichia coli and Salmonella typhimurium) to resist killing by antimicrobial peptides such as polymyxins. Members such as undecaprenyl phosphate-alpha-4-amino-4-deoxy-L-arabinose arabinosyl transferase are predicted to have 12 trans-membrane regions. The N-terminal portion of these proteins is hypothesized to have a conserved glycosylation activity which is shared between distantly related oligosaccharyltransferases ArnT and PglB families. |
| 406576 | PMT_4TMC | 2.52e-79 | 542 | 764 | 1 | 197 | C-terminal four TMM region of protein-O-mannosyltransferase. PMT_4TMC is the C-terminal four membrane-pass region of protein-O-mannosyltransferases and similar enzymes. |
| 397103 | MIR | 5.53e-20 | 352 | 500 | 13 | 170 | MIR domain. The MIR (protein mannosyltransferase, IP3R and RyR) domain is a domain that may have a ligand transferase function. |
| 197746 | MIR | 1.39e-08 | 332 | 390 | 2 | 56 | Domain in ryanodine and inositol trisphosphate receptors and protein O-mannosyltransferases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 772 | 1 | 768 | |
| 0.0 | 1 | 772 | 1 | 770 | |
| 0.0 | 1 | 772 | 1 | 770 | |
| 0.0 | 1 | 772 | 1 | 770 | |
| 0.0 | 1 | 772 | 1 | 770 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.31e-97 | 51 | 767 | 50 | 728 | Structure of S. cerevisiae protein O-mannosyltransferase Pmt1-Pmt2 complex bound to the sugar donor and a peptide acceptor [Saccharomyces cerevisiae W303],6P2R_A Structure of S. cerevisiae protein O-mannosyltransferase Pmt1-Pmt2 complex bound to the sugar donor [Saccharomyces cerevisiae W303] |
|
| 1.07e-89 | 8 | 681 | 20 | 696 | Structure of S. cerevisiae protein O-mannosyltransferase Pmt1-Pmt2 complex bound to the sugar donor and a peptide acceptor [Saccharomyces cerevisiae W303],6P2R_B Structure of S. cerevisiae protein O-mannosyltransferase Pmt1-Pmt2 complex bound to the sugar donor [Saccharomyces cerevisiae W303] |
|
| 7.21e-09 | 332 | 426 | 16 | 105 | Crystal structure of the SDF2-like protein from Arabidopsis thaliana [Arabidopsis thaliana],3MAL_B Crystal structure of the SDF2-like protein from Arabidopsis thaliana [Arabidopsis thaliana] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.19e-246 | 47 | 772 | 44 | 755 | Dolichyl-phosphate-mannose--protein mannosyltransferase 4 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=PMT4 PE=2 SV=2 |
|
| 2.58e-236 | 47 | 772 | 49 | 762 | Dolichyl-phosphate-mannose--protein mannosyltransferase 4 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=PMT4 PE=1 SV=1 |
|
| 3.39e-230 | 49 | 764 | 57 | 767 | Dolichyl-phosphate-mannose--protein mannosyltransferase 4 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=ogm4 PE=3 SV=1 |
|
| 1.33e-109 | 54 | 764 | 71 | 751 | Protein O-mannosyl-transferase 2 OS=Danio rerio OX=7955 GN=pomt2 PE=1 SV=1 |
|
| 6.53e-106 | 49 | 763 | 36 | 736 | Protein O-mannosyl-transferase 1 OS=Mus musculus OX=10090 GN=Pomt1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999882 | 0.000103 |
TMHMM Annotations download full data without filtering help
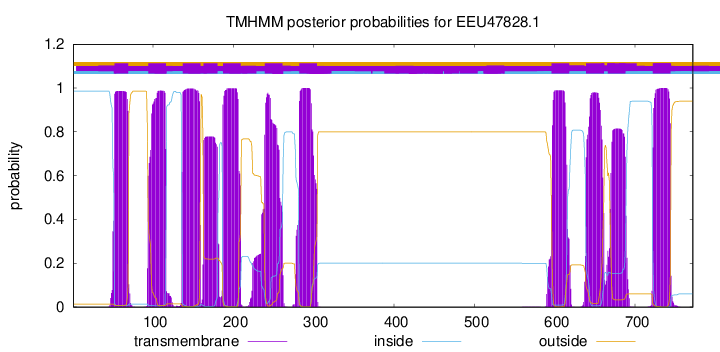
| Start | End |
|---|---|
| 52 | 69 |
| 94 | 116 |
| 137 | 159 |
| 163 | 180 |
| 187 | 209 |
| 239 | 261 |
| 282 | 304 |
| 596 | 618 |
| 639 | 661 |
| 666 | 688 |
| 722 | 744 |
