You are browsing environment: FUNGIDB
CAZyme Information: EEU45642.1
You are here: Home > Sequence: EEU45642.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium vanettenii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium vanettenii | |||||||||||
| CAZyme ID | EEU45642.1 | |||||||||||
| CAZy Family | PL3 | |||||||||||
| CAZyme Description | Glycoside hydrolase [Source:UniProtKB/TrEMBL;Acc:C7YSI0] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 227625 | Scw11 | 1.13e-28 | 6 | 261 | 40 | 267 | Exo-beta-1,3-glucanase, GH17 family [Carbohydrate transport and metabolism]. |
| 366033 | Glyco_hydro_17 | 7.77e-09 | 71 | 279 | 60 | 283 | Glycosyl hydrolases family 17. |
| 381626 | GH113_mannanase-like | 0.001 | 146 | 236 | 148 | 232 | Glycoside hydrolase family 113 beta-1,4-mannanase and similar proteins. Mannan endo-1,4-beta mannosidase (E.C 3.2.1.78) randomly cleaves (1->4)-beta-D-mannosidic linkages in mannans, galactomannans and glucomannans and is also called beta-1,4-mannanase, endo-1,4-beta-mannanase, endo-beta-1,4-mannase, beta-mannanase B, beta-1, 4-mannan 4-mannanohydrolase, endo-beta-mannanase, beta-D-mannanase, 1,4-beta-D-mannan mannanohydrolase, and 4-beta-D-mannan mannanohydrolase. (1->4)-beta-linked mannans are polysaccharides with a linear polymer backbone of (1->4)-beta-linked mannose units (in plants and fungi) or alternating mannose and glucose/galactose units (glucomannan in plants and fungi, and galactomannan and galactoglucomannan in plants), such as in the hemicellulose fraction of hard- and softwoods. Complete degradation of mannan requires a series of enzymes, including beta-1,4-mannanase. According to the CAZy database beta-1,4-mannanases are grouped into various glycoside hydrolase (GH) families; GH family 113 beta-1,4-mannanases include mostly bacterial and archaeal sequences. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.32e-225 | 1 | 308 | 11 | 318 | |
| 1.25e-198 | 1 | 308 | 11 | 318 | |
| 1.44e-198 | 1 | 308 | 11 | 318 | |
| 2.37e-197 | 1 | 308 | 11 | 318 | |
| 2.37e-197 | 1 | 308 | 11 | 318 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.33e-21 | 47 | 262 | 58 | 253 | Crystal structure of glycoside hydrolase family 17 beta-1,3-glucanosyltransferase from Rhizomucor miehei [Rhizomucor miehei CAU432] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.42e-103 | 1 | 302 | 10 | 311 | Probable glucan endo-1,3-beta-glucosidase eglC OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=eglC PE=3 SV=1 |
|
| 1.67e-99 | 1 | 302 | 10 | 310 | Probable glucan endo-1,3-beta-glucosidase eglC OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=eglC PE=3 SV=2 |
|
| 3.95e-98 | 1 | 302 | 10 | 310 | Probable glucan endo-1,3-beta-glucosidase eglC OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=eglC PE=3 SV=1 |
|
| 7.01e-98 | 1 | 302 | 10 | 310 | Probable glucan endo-1,3-beta-glucosidase eglC OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=eglC PE=3 SV=1 |
|
| 7.89e-98 | 1 | 302 | 10 | 310 | Probable glucan endo-1,3-beta-glucosidase eglC OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=eglC PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
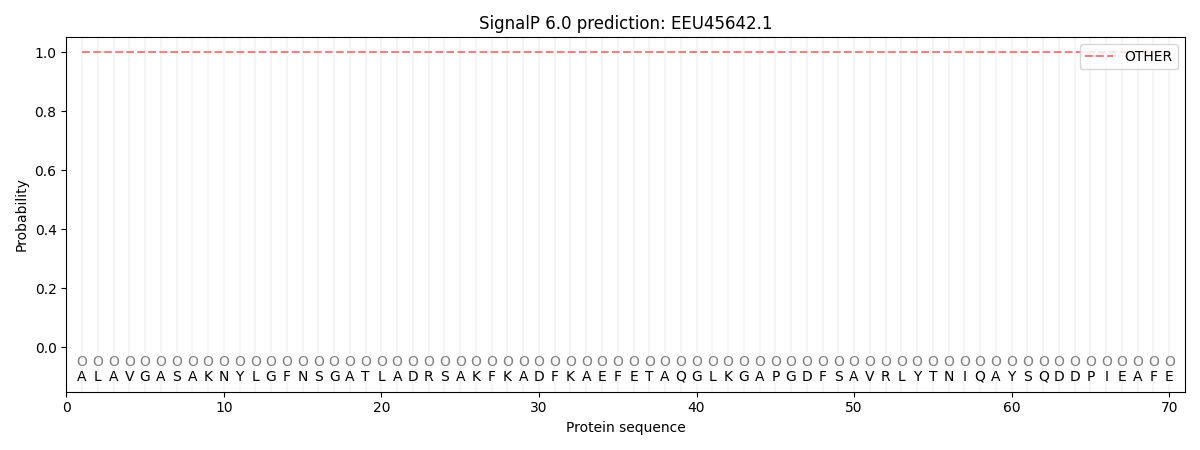
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000066 | 0.000002 |
