You are browsing environment: FUNGIDB
CAZyme Information: EEU45349.1
You are here: Home > Sequence: EEU45349.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
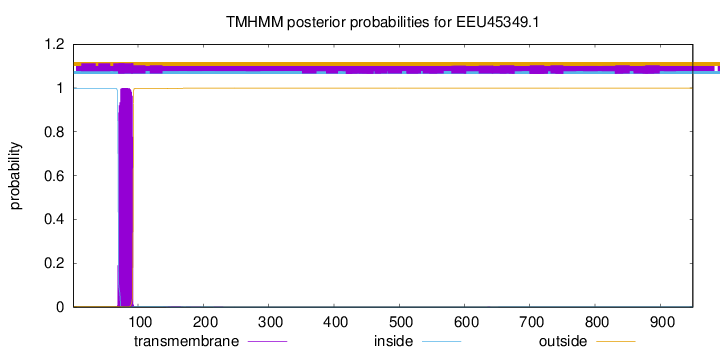
TMHMM annotations
Basic Information help
| Species | Fusarium vanettenii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium vanettenii | |||||||||||
| CAZyme ID | EEU45349.1 | |||||||||||
| CAZy Family | GT109 | |||||||||||
| CAZyme Description | glycoside hydrolase family 3 | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.21:13 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 161 | 374 | 8.1e-64 | 0.9722222222222222 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 224389 | BglX | 3.01e-57 | 171 | 489 | 67 | 351 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| 185053 | PRK15098 | 1.38e-52 | 104 | 944 | 16 | 762 | beta-glucosidase BglX. |
| 396478 | Glyco_hydro_3_C | 2.52e-51 | 477 | 725 | 1 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
| 395747 | Glyco_hydro_3 | 6.41e-35 | 170 | 412 | 74 | 313 | Glycosyl hydrolase family 3 N terminal domain. |
| 178629 | PLN03080 | 7.88e-33 | 75 | 902 | 5 | 740 | Probable beta-xylosidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 950 | 1 | 950 | |
| 0.0 | 1 | 944 | 1 | 936 | |
| 0.0 | 1 | 944 | 1 | 936 | |
| 0.0 | 1 | 944 | 1 | 936 | |
| 0.0 | 1 | 944 | 1 | 937 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.42e-262 | 104 | 940 | 4 | 831 | Chain A, Beta-glucosidase [Thermochaetoides thermophila] |
|
| 2.88e-258 | 104 | 940 | 7 | 834 | Three-dimensional structures of two heavily N-glycosylated Aspergillus sp. Family GH3 beta-D-glucosidases [Aspergillus oryzae],5FJJ_B Three-dimensional structures of two heavily N-glycosylated Aspergillus sp. Family GH3 beta-D-glucosidases [Aspergillus oryzae],5FJJ_C Three-dimensional structures of two heavily N-glycosylated Aspergillus sp. Family GH3 beta-D-glucosidases [Aspergillus oryzae],5FJJ_D Three-dimensional structures of two heavily N-glycosylated Aspergillus sp. Family GH3 beta-D-glucosidases [Aspergillus oryzae] |
|
| 3.08e-258 | 104 | 940 | 6 | 836 | Three-dimensional structures of two heavily N-glycosylated Aspergillus sp. Family GH3 beta-D-glucosidases [Aspergillus fumigatus],5FJI_B Three-dimensional structures of two heavily N-glycosylated Aspergillus sp. Family GH3 beta-D-glucosidases [Aspergillus fumigatus] |
|
| 1.63e-254 | 104 | 940 | 6 | 833 | Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus [Aspergillus aculeatus],4IIB_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus [Aspergillus aculeatus],4IIC_A Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with isofagomine [Aspergillus aculeatus],4IIC_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with isofagomine [Aspergillus aculeatus],4IID_A Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with 1-deoxynojirimycin [Aspergillus aculeatus],4IID_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with 1-deoxynojirimycin [Aspergillus aculeatus],4IIE_A Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with calystegine B(2) [Aspergillus aculeatus],4IIE_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with calystegine B(2) [Aspergillus aculeatus],4IIF_A Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with castanospermine [Aspergillus aculeatus],4IIF_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with castanospermine [Aspergillus aculeatus],4IIG_A Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with D-glucose [Aspergillus aculeatus],4IIG_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with D-glucose [Aspergillus aculeatus],4IIH_A Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with thiocellobiose [Aspergillus aculeatus],4IIH_B Crystal structure of beta-glucosidase 1 from Aspergillus aculeatus in complex with thiocellobiose [Aspergillus aculeatus] |
|
| 2.22e-253 | 104 | 940 | 25 | 847 | Chain A, Beta-glucosidase [Rasamsonia emersonii],5JU6_B Chain B, Beta-glucosidase [Rasamsonia emersonii],5JU6_C Chain C, Beta-glucosidase [Rasamsonia emersonii],5JU6_D Chain D, Beta-glucosidase [Rasamsonia emersonii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 86 | 940 | 136 | 1017 | Probable beta-glucosidase E OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=bglE PE=3 SV=1 |
|
| 0.0 | 72 | 940 | 174 | 1046 | Probable beta-glucosidase E OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=bglE PE=3 SV=1 |
|
| 0.0 | 72 | 944 | 162 | 1033 | Probable beta-glucosidase E OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=bglE PE=3 SV=1 |
|
| 0.0 | 72 | 944 | 162 | 1033 | Probable beta-glucosidase E OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=bglE PE=3 SV=1 |
|
| 0.0 | 72 | 944 | 164 | 1045 | Probable beta-glucosidase E OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=bglE PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
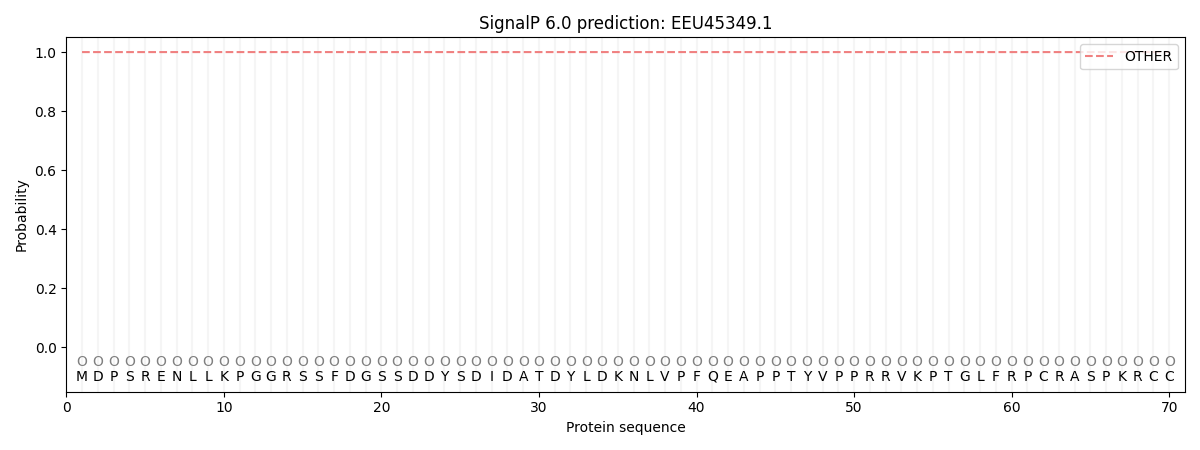
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000061 | 0.000000 |