You are browsing environment: FUNGIDB
CAZyme Information: EEU44048.1
You are here: Home > Sequence: EEU44048.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium vanettenii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium vanettenii | |||||||||||
| CAZyme ID | EEU44048.1 | |||||||||||
| CAZy Family | GH72 | |||||||||||
| CAZyme Description | alpha-1,2-Mannosidase [Source:UniProtKB/TrEMBL;Acc:C7YVS4] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.113:7 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH47 | 82 | 541 | 7.1e-159 | 0.9955156950672646 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396217 | Glyco_hydro_47 | 0.0 | 81 | 541 | 1 | 453 | Glycosyl hydrolase family 47. Members of this family are alpha-mannosidases that catalyze the hydrolysis of the terminal 1,2-linked alpha-D-mannose residues in the oligo-mannose oligosaccharide Man(9)(GlcNAc)(2). |
| 240427 | PTZ00470 | 1.20e-127 | 38 | 544 | 40 | 521 | glycoside hydrolase family 47 protein; Provisional |
| 224250 | YyaL | 8.14e-05 | 163 | 261 | 474 | 574 | Uncharacterized conserved protein YyaL, SSP411 family, contains thoiredoxin and six-hairpin glycosidase-like domains [General function prediction only]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 544 | 1 | 583 | |
| 4.19e-277 | 14 | 542 | 34 | 563 | |
| 1.35e-270 | 24 | 541 | 43 | 560 | |
| 1.48e-267 | 24 | 543 | 42 | 561 | |
| 4.22e-267 | 24 | 543 | 42 | 561 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.50e-80 | 74 | 538 | 5 | 448 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase and Man9GlcNAc2-PA complex [Homo sapiens] |
|
| 4.13e-80 | 74 | 538 | 10 | 453 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase [Homo sapiens] |
|
| 4.36e-80 | 74 | 538 | 10 | 453 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With 1-Deoxymannojirimycin [Homo sapiens],1FO3_A Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With Kifunensine [Homo sapiens] |
|
| 2.08e-79 | 74 | 538 | 5 | 448 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens],5KK7_B Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens] |
|
| 3.34e-79 | 74 | 538 | 88 | 531 | Crystal Structure Of Human Class I alpha-1,2-Mannosidase In Complex With Thio-Disaccharide Substrate Analogue [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.27e-77 | 74 | 538 | 249 | 692 | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase OS=Homo sapiens OX=9606 GN=MAN1B1 PE=1 SV=2 |
|
| 4.23e-71 | 72 | 540 | 95 | 535 | Mannosyl-oligosaccharide 1,2-alpha-mannosidase MNS1 OS=Arabidopsis thaliana OX=3702 GN=MNS1 PE=1 SV=1 |
|
| 4.71e-69 | 74 | 538 | 128 | 617 | Mannosyl-oligosaccharide 1,2-alpha-mannosidase MNS3 OS=Arabidopsis thaliana OX=3702 GN=MNS3 PE=1 SV=1 |
|
| 9.00e-69 | 73 | 538 | 206 | 650 | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase OS=Rattus norvegicus OX=10116 GN=Man1b1 PE=2 SV=2 |
|
| 1.78e-68 | 73 | 538 | 207 | 651 | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase OS=Mus musculus OX=10090 GN=Man1b1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
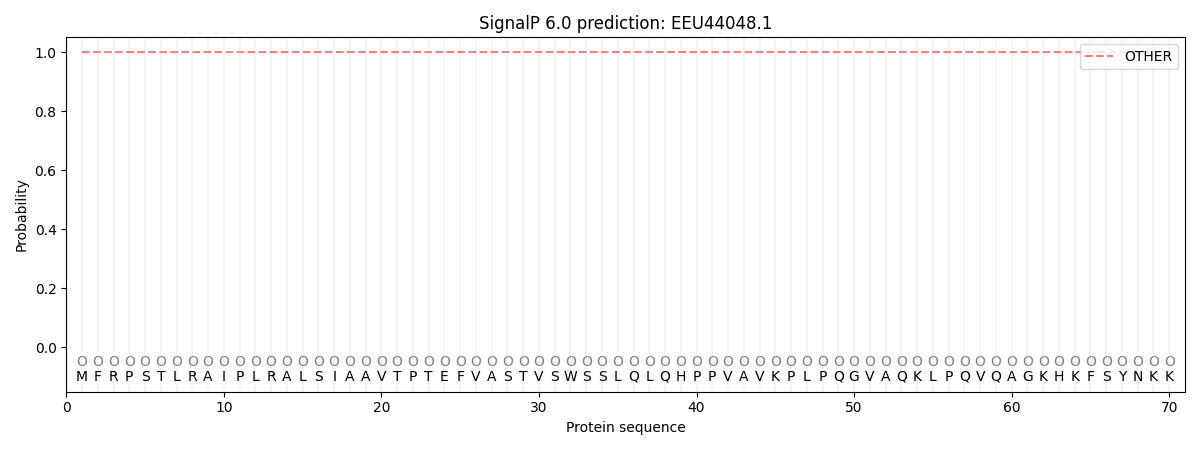
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000053 | 0.000000 |
