You are browsing environment: FUNGIDB
CAZyme Information: EEU37841.1
You are here: Home > Sequence: EEU37841.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Fusarium vanettenii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Nectriaceae; Fusarium; Fusarium vanettenii | |||||||||||
| CAZyme ID | EEU37841.1 | |||||||||||
| CAZy Family | GH13 | |||||||||||
| CAZyme Description | Oxidoreductase [Source:UniProtKB/TrEMBL;Acc:C7ZDK3] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH109 | 9 | 171 | 7.4e-28 | 0.40601503759398494 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 223745 | MviM | 6.36e-42 | 9 | 224 | 5 | 213 | Predicted dehydrogenase [General function prediction only]. |
| 397161 | GFO_IDH_MocA_C | 8.26e-21 | 148 | 435 | 1 | 204 | Oxidoreductase family, C-terminal alpha/beta domain. This family of enzymes utilize NADP or NAD. This family is called the GFO/IDH/MOCA family in swiss-prot. |
| 396129 | GFO_IDH_MocA | 2.18e-17 | 9 | 136 | 2 | 120 | Oxidoreductase family, NAD-binding Rossmann fold. This family of enzymes utilize NADP or NAD. This family is called the GFO/IDH/MOCA family in swiss-prot. |
| 182305 | PRK10206 | 1.31e-05 | 60 | 166 | 42 | 154 | putative oxidoreductase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 6.54e-25 | 9 | 437 | 396 | 753 | |
| 3.01e-15 | 3 | 430 | 13 | 393 | |
| 1.14e-14 | 39 | 286 | 33 | 268 | |
| 5.99e-11 | 2 | 427 | 49 | 431 | |
| 5.99e-11 | 2 | 427 | 49 | 431 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.12e-12 | 9 | 219 | 4 | 198 | Crystal structure of probable dehydrogenase TM_0414 from Thermotoga maritima [Thermotoga maritima],3EZY_B Crystal structure of probable dehydrogenase TM_0414 from Thermotoga maritima [Thermotoga maritima],3EZY_C Crystal structure of probable dehydrogenase TM_0414 from Thermotoga maritima [Thermotoga maritima],3EZY_D Crystal structure of probable dehydrogenase TM_0414 from Thermotoga maritima [Thermotoga maritima] |
|
| 1.17e-11 | 34 | 213 | 28 | 191 | Crystal structure of a putative myo-inositol dehydrogenase from Sinorhizobium meliloti 1021 (Target PSI-012312) [Sinorhizobium meliloti 1021],4HKT_B Crystal structure of a putative myo-inositol dehydrogenase from Sinorhizobium meliloti 1021 (Target PSI-012312) [Sinorhizobium meliloti 1021],4HKT_C Crystal structure of a putative myo-inositol dehydrogenase from Sinorhizobium meliloti 1021 (Target PSI-012312) [Sinorhizobium meliloti 1021],4HKT_D Crystal structure of a putative myo-inositol dehydrogenase from Sinorhizobium meliloti 1021 (Target PSI-012312) [Sinorhizobium meliloti 1021] |
|
| 1.47e-11 | 9 | 214 | 32 | 227 | Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_B Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_C Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_D Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_E Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_F Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_G Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_H Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_I Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_J Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_K Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_L Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_M Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_N Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_O Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I],3Q2K_P Crystal structure of the WlbA dehydrogenase from Bordetella pertussis in complex with NADH and UDP-GlcNAcA [Bordetella pertussis Tohama I] |
|
| 2.83e-11 | 9 | 202 | 4 | 186 | The crystal structure of the putative dehydrogenase from Bordetella bronchiseptica RB50 [Bordetella bronchiseptica] |
|
| 1.01e-10 | 55 | 210 | 46 | 201 | Chain A, GLUCOSE-FRUCTOSE OXIDOREDUCTASE [Zymomonas mobilis],1EVJ_B Chain B, GLUCOSE-FRUCTOSE OXIDOREDUCTASE [Zymomonas mobilis],1EVJ_C Chain C, GLUCOSE-FRUCTOSE OXIDOREDUCTASE [Zymomonas mobilis],1EVJ_D Chain D, GLUCOSE-FRUCTOSE OXIDOREDUCTASE [Zymomonas mobilis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.59e-75 | 11 | 438 | 5 | 422 | Putative oxidoreductase YteT OS=Bacillus subtilis (strain 168) OX=224308 GN=yteT PE=2 SV=1 |
|
| 1.06e-11 | 2 | 427 | 49 | 431 | Glycosyl hydrolase family 109 protein OS=Shewanella baltica (strain OS155 / ATCC BAA-1091) OX=325240 GN=Sbal_2793 PE=3 SV=1 |
|
| 1.88e-11 | 2 | 427 | 49 | 431 | Glycosyl hydrolase family 109 protein OS=Shewanella baltica (strain OS185) OX=402882 GN=Shew185_2813 PE=3 SV=1 |
|
| 2.49e-11 | 34 | 213 | 27 | 190 | Inositol 2-dehydrogenase OS=Rhizobium meliloti (strain 1021) OX=266834 GN=idhA PE=1 SV=2 |
|
| 3.33e-11 | 2 | 427 | 49 | 431 | Glycosyl hydrolase family 109 protein OS=Shewanella baltica (strain OS195) OX=399599 GN=Sbal195_2933 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
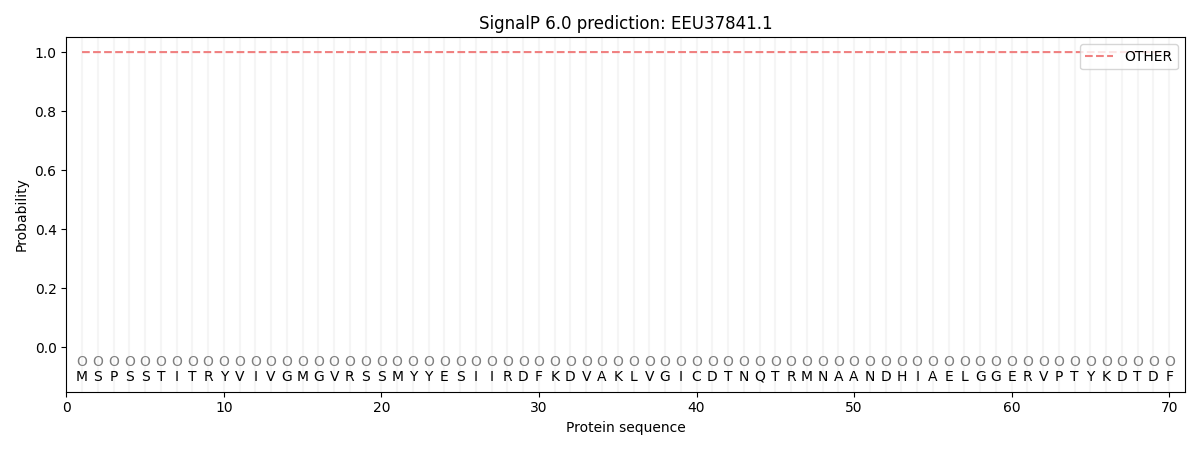
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000058 | 0.000000 |
