You are browsing environment: FUNGIDB
CAZyme Information: EEP79928.1
You are here: Home > Sequence: EEP79928.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
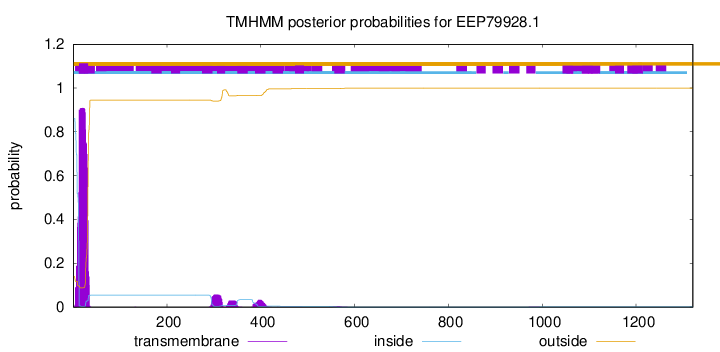
TMHMM annotations
Basic Information help
| Species | Uncinocarpus reesii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Onygenaceae; Uncinocarpus; Uncinocarpus reesii | |||||||||||
| CAZyme ID | EEP79928.1 | |||||||||||
| CAZy Family | GH81 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.58:16 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH55 | 534 | 891 | 1.2e-142 | 0.4756756756756757 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 403800 | Pectate_lyase_3 | 1.31e-77 | 257 | 451 | 1 | 212 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| 403800 | Pectate_lyase_3 | 2.93e-07 | 534 | 587 | 7 | 62 | Pectate lyase superfamily protein. This family of proteins possesses a beta helical structure like Pectate lyase. This family is most closely related to glycosyl hydrolase family 28. |
| 227721 | Pgu1 | 4.16e-05 | 259 | 309 | 84 | 125 | Polygalacturonase [Carbohydrate transport and metabolism]. |
| 396179 | LysM | 1.56e-04 | 1097 | 1142 | 1 | 43 | LysM domain. The LysM (lysin motif) domain is about 40 residues long. It is found in a variety of enzymes involved in bacterial cell wall degradation. This domain may have a general peptidoglycan binding function. The structure of this domain is known. |
| 314993 | End_N_terminal | 1.83e-04 | 265 | 329 | 1 | 61 | N terminal extension of bacteriophage endosialidase. This domain family is found in bacteria and viruses, and is approximately 70 amino acids in length. This domain is found in the bacteriophage protein endosialidase. The two N-terminal domains (this domain and the beta propeller) assemble in the compact 'cap' whereas the C-terminal domain forms an extended tail-like structure. The very N-terminal part of the 'cap' region (residues 246 to 312) holds the only alpha-helix of the protein and is presumably the residual part of the deleted N-terminal head-binding domain. The endosialidase protein complexes to form homotrimeric molecules. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 11 | 1302 | 11 | 1371 | |
| 0.0 | 85 | 1304 | 1 | 1312 | |
| 0.0 | 23 | 1308 | 7 | 1369 | |
| 0.0 | 17 | 1302 | 10 | 1369 | |
| 0.0 | 51 | 1301 | 32 | 1351 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.66e-179 | 236 | 891 | 6 | 752 | Chain A, Beta-1,3-glucanase [Thermochaetoides thermophila],5M60_A Chain A, Beta-1,3-glucanase [Thermochaetoides thermophila] |
|
| 1.49e-169 | 236 | 894 | 26 | 758 | Chain A, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQN_B Chain B, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQO_A Chain A, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium],3EQO_B Chain B, Glucan 1,3-beta-glucosidase [Phanerodontia chrysosporium] |
|
| 5.12e-07 | 259 | 324 | 76 | 139 | Chain A, Putative pectin lyase [Geobacillus virus E2],7CHU_B Chain B, Putative pectin lyase [Geobacillus virus E2],7CHU_C Chain C, Putative pectin lyase [Geobacillus virus E2] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 5.04e-168 | 236 | 888 | 43 | 870 | Probable glucan endo-1,3-beta-glucosidase ARB_02077 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_02077 PE=1 SV=1 |
|
| 9.99e-134 | 219 | 888 | 29 | 781 | Glucan 1,3-beta-glucosidase OS=Cochliobolus carbonum OX=5017 GN=EXG1 PE=1 SV=1 |
|
| 2.23e-38 | 537 | 841 | 401 | 705 | Glucan endo-1,3-beta-glucosidase BGN13.1 OS=Trichoderma harzianum OX=5544 GN=bgn13.1 PE=1 SV=1 |
|
| 2.66e-08 | 914 | 1298 | 30 | 422 | LysM domain-containing protein ARB_01155/01156 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_01155/01156 PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000363 | 0.999614 | CS pos: 31-32. Pr: 0.9738 |