You are browsing environment: FUNGIDB
CAZyme Information: EDK43323.1
You are here: Home > Sequence: EDK43323.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Lodderomyces elongisporus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Saccharomycetes; ; Debaryomycetaceae; Lodderomyces; Lodderomyces elongisporus | |||||||||||
| CAZyme ID | EDK43323.1 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | glucan 1,3-beta-glucosidase precursor | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.58:29 | 3.2.1.21:6 | 3.2.1.75:2 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 104 | 394 | 1.1e-125 | 0.9966996699669967 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 225344 | BglC | 8.79e-79 | 54 | 424 | 19 | 395 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| 395098 | Cellulase | 7.78e-08 | 109 | 260 | 24 | 165 | Cellulase (glycosyl hydrolase family 5). |
| 396011 | AP_endonuc_2 | 0.001 | 121 | 225 | 79 | 179 | Xylose isomerase-like TIM barrel. This TIM alpha/beta barrel structure is found in xylose isomerase and in endonuclease IV (EC:3.1.21.2). This domain is also found in the N termini of bacterial myo-inositol catabolism proteins. These are involved in the myo-inositol catabolism pathway, and is required for growth on myo-inositol in Rhizobium leguminosarum bv. viciae. |
| 277321 | MPP_ASMase | 0.004 | 102 | 222 | 141 | 251 | acid sphingomyelinase and related proteins, metallophosphatase domain. Acid sphingomyelinase (ASMase) is a ubiquitously expressed phosphodiesterase which hydrolyzes sphingomyelin in acid pH conditions to form ceramide, a bioactive second messenger, as part of the sphingomyelin signaling pathway. ASMase is localized at the noncytosolic leaflet of biomembranes (for example the luminal leaflet of endosomes, lysosomes and phagosomes, and the extracellular leaflet of plasma membranes). ASMase-deficient humans develop Niemann-Pick disease. This disease is characterized by lysosomal storage of sphingomyelin in all tissues. Although ASMase-deficient mice are resistant to stress-induced apoptosis, they have greater susceptibility to bacterial infection. The latter correlates with defective phagolysosomal fusion and antibacterial killing activity in ASMase-deficient macrophages. ASMase belongs to the metallophosphatase (MPP) superfamily. MPPs are functionally diverse, but all share a conserved domain with an active site consisting of two metal ions (usually manganese, iron, or zinc) coordinated with octahedral geometry by a cage of histidine, aspartate, and asparagine residues. The MPP superfamily includes: the phosphoprotein phosphatases (PPPs), Mre11/SbcD-like exonucleases, Dbr1-like RNA lariat debranching enzymes, YfcE-like phosphodiesterases, purple acid phosphatases (PAPs), YbbF-like UDP-2,3-diacylglucosamine hydrolases, and acid sphingomyelinases (ASMases). The conserved domain is a double beta-sheet sandwich with a di-metal active site made up of residues located at the C-terminal side of the sheets. This domain is thought to allow for productive metal coordination. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.14e-236 | 6 | 425 | 4 | 421 | |
| 3.39e-235 | 7 | 427 | 5 | 423 | |
| 1.03e-226 | 36 | 426 | 24 | 431 | |
| 1.08e-225 | 36 | 426 | 31 | 438 | |
| 1.08e-225 | 36 | 426 | 31 | 438 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.13e-227 | 45 | 426 | 1 | 394 | Exo-b-(1,3)-glucanase From Candida Albicans At 1.85 A Resolution [Candida albicans],1EQC_A Exo-b-(1,3)-glucanase From Candida Albicans In Complex With Castanospermine At 1.85 A [Candida albicans] |
|
| 2.66e-227 | 45 | 426 | 7 | 400 | Exo-B-(1,3)-Glucanase from Candida Albicans in complex with unhydrolysed and covalently linked 2,4-dinitrophenyl-2-deoxy-2-fluoro-B-D-glucopyranoside at 1.9 A [Candida albicans] |
|
| 7.62e-227 | 45 | 426 | 7 | 400 | Chain A, Hypothetical protein XOG1 [Candida albicans] |
|
| 7.62e-227 | 45 | 426 | 7 | 400 | Chain A, Hypothetical protein XOG1 [Candida albicans] |
|
| 1.48e-226 | 45 | 426 | 6 | 399 | The structure of E292S glycosynthase variant of exo-1,3-beta-glucanase from Candida albicans at 1.85A resolution [Candida albicans SC5314],4M81_A The structure of E292S glycosynthase variant of exo-1,3-beta-glucanase from Candida albicans complexed with 1-fluoro-alpha-D-glucopyranoside (donor) and p-nitrophenyl beta-D-glucopyranoside (acceptor) at 1.86A resolution [Candida albicans SC5314],4M82_A The structure of E292S glycosynthase variant of exo-1,3-beta-glucanase from Candida albicans complexed with p-nitrophenyl-gentiobioside (product) at 1.6A resolution [Candida albicans SC5314] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.88e-226 | 36 | 426 | 31 | 438 | Glucan 1,3-beta-glucosidase OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=XOG1 PE=1 SV=5 |
|
| 9.20e-210 | 31 | 425 | 19 | 425 | Glucan 1,3-beta-glucosidase OS=Schwanniomyces occidentalis OX=27300 PE=3 SV=1 |
|
| 3.68e-202 | 39 | 425 | 23 | 425 | Glucan 1,3-beta-glucosidase OS=Candida oleophila OX=45573 GN=EXG1 PE=3 SV=1 |
|
| 1.36e-195 | 46 | 424 | 35 | 427 | Glucan 1,3-beta-glucosidase 2 OS=Wickerhamomyces anomalus OX=4927 GN=EXG2 PE=3 SV=1 |
|
| 9.20e-180 | 40 | 425 | 35 | 439 | Glucan 1,3-beta-glucosidase OS=Lachancea kluyveri (strain ATCC 58438 / CBS 3082 / BCRC 21498 / NBRC 1685 / JCM 7257 / NCYC 543 / NRRL Y-12651) OX=226302 GN=EXG1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
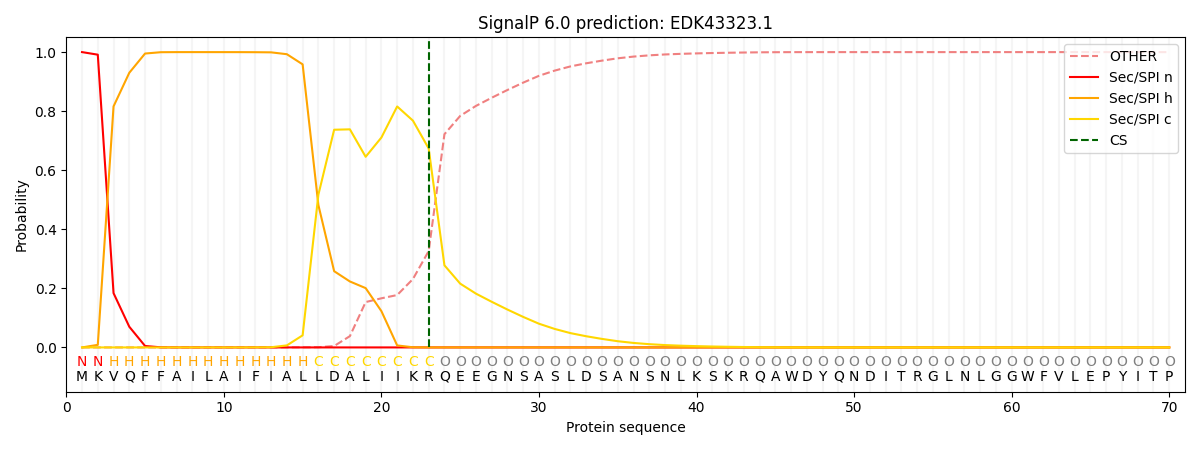
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000467 | 0.999514 | CS pos: 23-24. Pr: 0.6736 |
