You are browsing environment: FUNGIDB
CAZyme Information: EAW24240.1
You are here: Home > Sequence: EAW24240.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus fischeri | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus fischeri | |||||||||||
| CAZyme ID | EAW24240.1 | |||||||||||
| CAZy Family | PL4 | |||||||||||
| CAZyme Description | LysM domain protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 212030 | LysM | 1.15e-09 | 131 | 175 | 1 | 45 | Lysin Motif is a small domain involved in binding peptidoglycan. LysM, a small globular domain with approximately 40 amino acids, is a widespread protein module involved in binding peptidoglycan in bacteria and chitin in eukaryotes. The domain was originally identified in enzymes that degrade bacterial cell walls, but proteins involved in many other biological functions also contain this domain. It has been reported that the LysM domain functions as a signal for specific plant-bacteria recognition in bacterial pathogenesis. Many of these enzymes are modular and are composed of catalytic units linked to one or several repeats of LysM domains. LysM domains are found in bacteria and eukaryotes. |
| 212030 | LysM | 4.35e-08 | 226 | 270 | 1 | 45 | Lysin Motif is a small domain involved in binding peptidoglycan. LysM, a small globular domain with approximately 40 amino acids, is a widespread protein module involved in binding peptidoglycan in bacteria and chitin in eukaryotes. The domain was originally identified in enzymes that degrade bacterial cell walls, but proteins involved in many other biological functions also contain this domain. It has been reported that the LysM domain functions as a signal for specific plant-bacteria recognition in bacterial pathogenesis. Many of these enzymes are modular and are composed of catalytic units linked to one or several repeats of LysM domains. LysM domains are found in bacteria and eukaryotes. |
| 212030 | LysM | 1.60e-07 | 310 | 354 | 1 | 45 | Lysin Motif is a small domain involved in binding peptidoglycan. LysM, a small globular domain with approximately 40 amino acids, is a widespread protein module involved in binding peptidoglycan in bacteria and chitin in eukaryotes. The domain was originally identified in enzymes that degrade bacterial cell walls, but proteins involved in many other biological functions also contain this domain. It has been reported that the LysM domain functions as a signal for specific plant-bacteria recognition in bacterial pathogenesis. Many of these enzymes are modular and are composed of catalytic units linked to one or several repeats of LysM domains. LysM domains are found in bacteria and eukaryotes. |
| 197609 | LysM | 1.03e-06 | 132 | 177 | 1 | 44 | Lysin motif. |
| 396179 | LysM | 6.76e-05 | 133 | 159 | 1 | 27 | LysM domain. The LysM (lysin motif) domain is about 40 residues long. It is found in a variety of enzymes involved in bacterial cell wall degradation. This domain may have a general peptidoglycan binding function. The structure of this domain is known. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.39e-144 | 2 | 358 | 1 | 355 | |
| 1.25e-128 | 2 | 358 | 1 | 342 | |
| 2.29e-99 | 16 | 358 | 15 | 335 | |
| 5.18e-96 | 22 | 320 | 24 | 314 | |
| 4.46e-89 | 25 | 358 | 121 | 450 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.65e-80 | 30 | 357 | 149 | 456 | LysM domain-containing protein ARB_05157 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_05157 PE=1 SV=1 |
|
| 3.48e-19 | 121 | 179 | 379 | 437 | Probable peptidoglycan-N-acetylglucosamine deacetylase ARB_03699 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_03699 PE=1 SV=2 |
|
| 7.32e-15 | 36 | 182 | 29 | 184 | LysM domain-containing protein ARB_03438 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_03438 PE=3 SV=1 |
|
| 2.87e-12 | 24 | 179 | 356 | 536 | LysM domain-containing protein ARB_00327 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_00327 PE=3 SV=2 |
|
| 1.60e-07 | 118 | 177 | 360 | 419 | LysM domain-containing protein ARB_01155/01156 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_01155/01156 PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
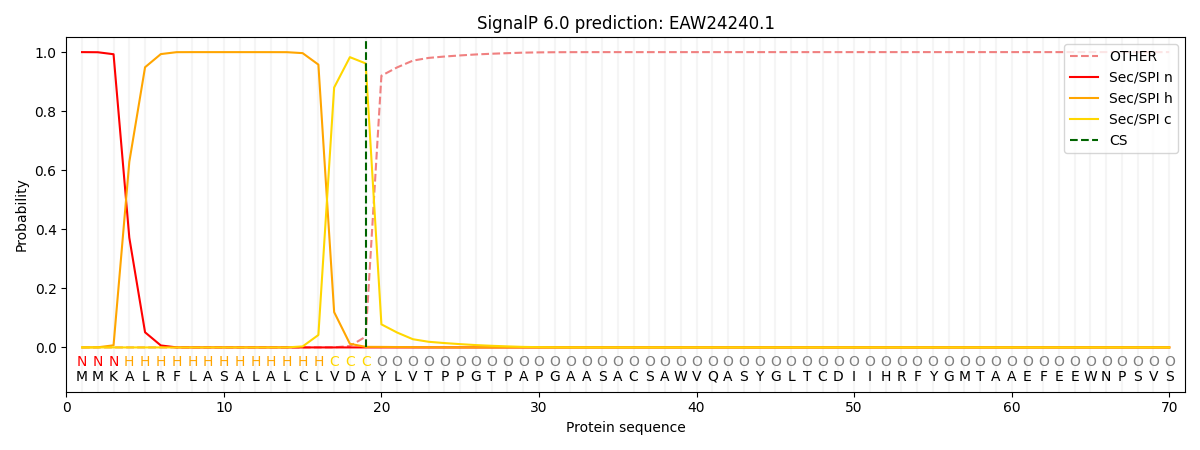
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000208 | 0.999741 | CS pos: 19-20. Pr: 0.9617 |
