You are browsing environment: FUNGIDB
CAZyme Information: EAW16482.1
You are here: Home > Sequence: EAW16482.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
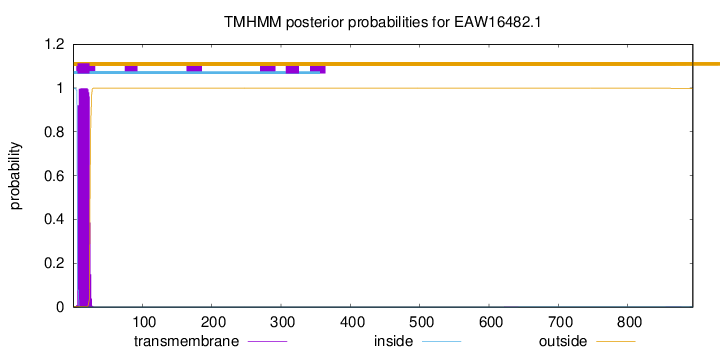
TMHMM annotations
Basic Information help
| Species | Aspergillus fischeri | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus fischeri | |||||||||||
| CAZyme ID | EAW16482.1 | |||||||||||
| CAZy Family | AA7 | |||||||||||
| CAZyme Description | class I alpha-mannosidase 1A | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.113:7 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH47 | 214 | 891 | 8e-176 | 0.9977578475336323 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396217 | Glyco_hydro_47 | 0.0 | 214 | 891 | 1 | 453 | Glycosyl hydrolase family 47. Members of this family are alpha-mannosidases that catalyze the hydrolysis of the terminal 1,2-linked alpha-D-mannose residues in the oligo-mannose oligosaccharide Man(9)(GlcNAc)(2). |
| 240427 | PTZ00470 | 3.05e-100 | 139 | 892 | 2 | 519 | glycoside hydrolase family 47 protein; Provisional |
| 411474 | fibronec_FbpA | 2.36e-04 | 36 | 122 | 194 | 275 | LPXTG-anchored fibronectin-binding protein FbpA. FbpA, a fibronectin-binding protein described in Streptococcus pyogenes, has a YSIRK-type (crosswall-targeting) signal peptide and a C-terminal LPXTG motif for covalent attachment to the cell wall. It is unrelated to the PavA-like protein from Streptococcus gordonii (see BlastRule NBR009716) that was given the identical name, so the phase LPXTG-anchored is added to the protein name for clarity. |
| 223021 | PHA03247 | 0.002 | 32 | 107 | 2883 | 2956 | large tegument protein UL36; Provisional |
| 237865 | PRK14951 | 0.003 | 24 | 95 | 409 | 478 | DNA polymerase III subunits gamma and tau; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 892 | 1 | 868 | |
| 0.0 | 1 | 892 | 1 | 869 | |
| 0.0 | 1 | 892 | 1 | 869 | |
| 0.0 | 1 | 894 | 1 | 851 | |
| 0.0 | 1 | 894 | 1 | 851 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.63e-49 | 215 | 889 | 13 | 449 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase and Man9GlcNAc2-PA complex [Homo sapiens] |
|
| 7.53e-49 | 215 | 889 | 18 | 454 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase [Homo sapiens] |
|
| 7.86e-49 | 215 | 889 | 18 | 454 | Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With 1-Deoxymannojirimycin [Homo sapiens],1FO3_A Crystal Structure Of Human Class I Alpha1,2-Mannosidase In Complex With Kifunensine [Homo sapiens] |
|
| 3.30e-48 | 215 | 889 | 13 | 449 | Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens],5KK7_B Crystal structure of the class I human endoplasmic reticulum 1,2-alpha-mannosidase T688A mutant and Thio-disaccharide substrate analog complex [Homo sapiens] |
|
| 3.56e-48 | 215 | 889 | 96 | 532 | Crystal Structure Of Human Class I alpha-1,2-Mannosidase In Complex With Thio-Disaccharide Substrate Analogue [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.86e-48 | 207 | 894 | 197 | 651 | Mannosyl-oligosaccharide alpha-1,2-mannosidase IA OS=Spodoptera frugiperda OX=7108 PE=1 SV=1 |
|
| 3.40e-47 | 154 | 889 | 160 | 652 | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase OS=Mus musculus OX=10090 GN=Man1b1 PE=1 SV=1 |
|
| 1.11e-46 | 206 | 889 | 206 | 651 | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase OS=Rattus norvegicus OX=10116 GN=Man1b1 PE=2 SV=2 |
|
| 1.83e-46 | 215 | 889 | 257 | 693 | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase OS=Homo sapiens OX=9606 GN=MAN1B1 PE=1 SV=2 |
|
| 4.09e-41 | 193 | 890 | 25 | 506 | Probable mannosyl-oligosaccharide alpha-1,2-mannosidase 1B OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=mns1B PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
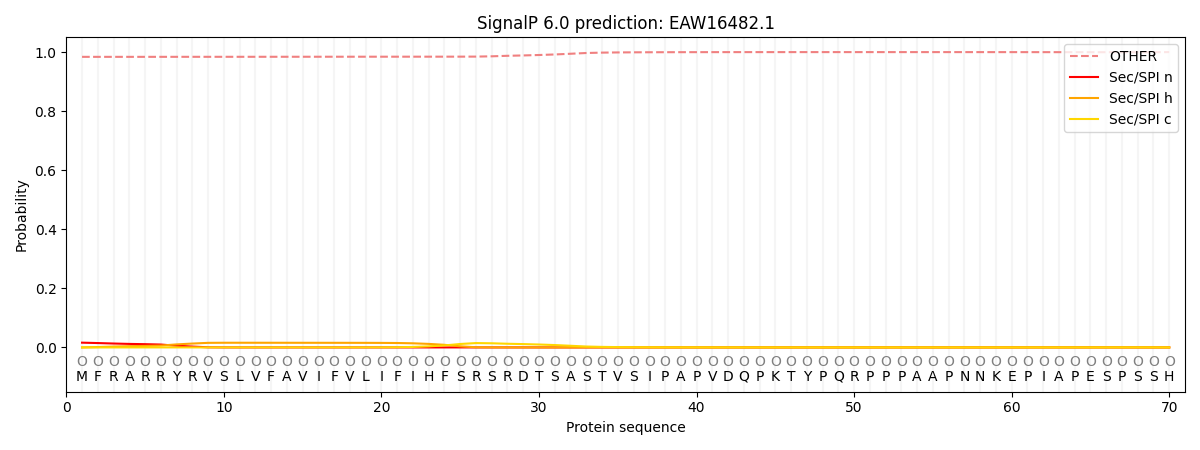
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.984803 | 0.015213 |