You are browsing environment: FUNGIDB
CAZyme Information: EAQ93832.1
You are here: Home > Sequence: EAQ93832.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Chaetomium globosum | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Sordariomycetes; ; Chaetomiaceae; Chaetomium; Chaetomium globosum | |||||||||||
| CAZyme ID | EAQ93832.1 | |||||||||||
| CAZy Family | GT76 | |||||||||||
| CAZyme Description | Endo-1,4-beta-xylanase [Source:UniProtKB/TrEMBL;Acc:Q2HCI7] | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.8:18 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH11 | 55 | 216 | 2.5e-57 | 0.9887005649717514 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395367 | Glyco_hydro_11 | 9.94e-93 | 54 | 214 | 1 | 175 | Glycosyl hydrolases family 11. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 8.73e-136 | 1 | 235 | 1 | 250 | |
| 3.43e-135 | 1 | 232 | 1 | 247 | |
| 3.21e-122 | 7 | 238 | 10 | 253 | |
| 9.19e-122 | 7 | 238 | 10 | 253 | |
| 9.19e-122 | 7 | 238 | 10 | 253 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.12e-69 | 51 | 221 | 12 | 194 | Chain A, Endo-1,4-beta-xylanase [Actinomycetia bacterium],7EO6_B Chain B, Endo-1,4-beta-xylanase [Actinomycetia bacterium] |
|
| 2.81e-69 | 51 | 218 | 9 | 191 | Chain A, Endo-1,4-beta-xylanase [Thermochaetoides thermophila],1H1A_B Chain B, Endo-1,4-beta-xylanase [Thermochaetoides thermophila] |
|
| 3.30e-69 | 51 | 218 | 9 | 191 | Chain A, endoxylanase 11A [Thermochaetoides thermophila],1XNK_B Chain B, endoxylanase 11A [Thermochaetoides thermophila] |
|
| 7.50e-69 | 49 | 217 | 5 | 188 | Chain A, Endo-1,4-beta-xylanase 2 [Trichoderma reesei],4XQD_B Chain B, Endo-1,4-beta-xylanase 2 [Trichoderma reesei],5ZF3_A Crystal Structures of Endo-beta-1,4-xylanase II Complexed with Xylotriose [Trichoderma reesei RUT C-30],5ZH0_A Crystal Structures of Endo-beta-1,4-xylanase II [Trichoderma reesei RUT C-30],5ZO0_A Neutron structure of xylanase at pD5.4 [Trichoderma reesei RUT C-30] |
|
| 7.74e-69 | 49 | 217 | 6 | 189 | Chain A, ENDO-1,4-BETA-XYLANASE II [Trichoderma reesei],1ENX_B Chain B, ENDO-1,4-BETA-XYLANASE II [Trichoderma reesei],1RED_A Chain A, ENDO-1,4-BETA-XYLANASE II [Trichoderma reesei],1RED_B Chain B, ENDO-1,4-BETA-XYLANASE II [Trichoderma reesei],1REE_A Chain A, ENDO-1,4-BETA-XYLANASE II [Trichoderma reesei],1REE_B Chain B, ENDO-1,4-BETA-XYLANASE II [Trichoderma reesei],1REF_A Chain A, ENDO-1,4-BETA-XYLANASE II [Trichoderma reesei],1REF_B Chain B, ENDO-1,4-BETA-XYLANASE II [Trichoderma reesei],1XYO_A Chain A, ENDO-1,4-BETA-XYLANASE II [Trichoderma reesei],1XYO_B Chain B, ENDO-1,4-BETA-XYLANASE II [Trichoderma reesei],1XYP_A Chain A, ENDO-1,4-BETA-XYLANASE II [Trichoderma reesei],1XYP_B Chain B, ENDO-1,4-BETA-XYLANASE II [Trichoderma reesei],2DFB_A Chain A, Endo-1,4-beta-xylanase 2 [Trichoderma reesei],2DFC_A Chain A, Endo-1,4-beta-xylanase 2 [Trichoderma reesei],3AKP_A Crystal structure of xylanase from Trichoderma longibrachiatum [Trichoderma longibrachiatum],3AKP_B Crystal structure of xylanase from Trichoderma longibrachiatum [Trichoderma longibrachiatum],3AKQ_A Crystal structure of xylanase from Trichoderma longibrachiatum [Trichoderma longibrachiatum],3AKR_A Crystal structure of xylanase from Trichoderma longibrachiatum [Trichoderma longibrachiatum],3AKS_A Crystal structure of xylanase from Trichoderma longibrachiatum [Trichoderma longibrachiatum],3AKT_A Crystal structure of xylanase from Trichoderma longibrachiatum [Trichoderma longibrachiatum],3AKT_B Crystal structure of xylanase from Trichoderma longibrachiatum [Trichoderma longibrachiatum],3LGR_A Chain A, Endo-1,4-beta-xylanase 2 [Trichoderma reesei],4HKW_A Chain A, Endo-1,4-beta-xylanase 2 [Trichoderma reesei],4S2D_A Chain A, Endo-1,4-beta-xylanase 2 [Trichoderma reesei],4S2F_A Chain A, Endo-1,4-beta-xylanase 2 [Trichoderma reesei],4S2G_A Chain A, Endo-1,4-beta-xylanase 2 [Trichoderma reesei],4S2H_A Chain A, Endo-1,4-beta-xylanase 2 [Trichoderma reesei],7MGU_A Chain A, Endo-1,4-beta-xylanase [Trichoderma reesei] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.03e-76 | 46 | 217 | 39 | 224 | Endo-1,4-beta-xylanase A OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=xlnA PE=1 SV=1 |
|
| 3.61e-75 | 8 | 217 | 12 | 227 | Probable endo-1,4-beta-xylanase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=xlnA PE=3 SV=1 |
|
| 5.82e-75 | 42 | 217 | 42 | 231 | Probable endo-1,4-beta-xylanase A OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=xlnA PE=3 SV=2 |
|
| 5.82e-75 | 42 | 217 | 42 | 231 | Endo-1,4-beta-xylanase A OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=xlnA PE=1 SV=1 |
|
| 7.99e-75 | 46 | 217 | 45 | 230 | Probable endo-1,4-beta-xylanase A OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=xlnA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
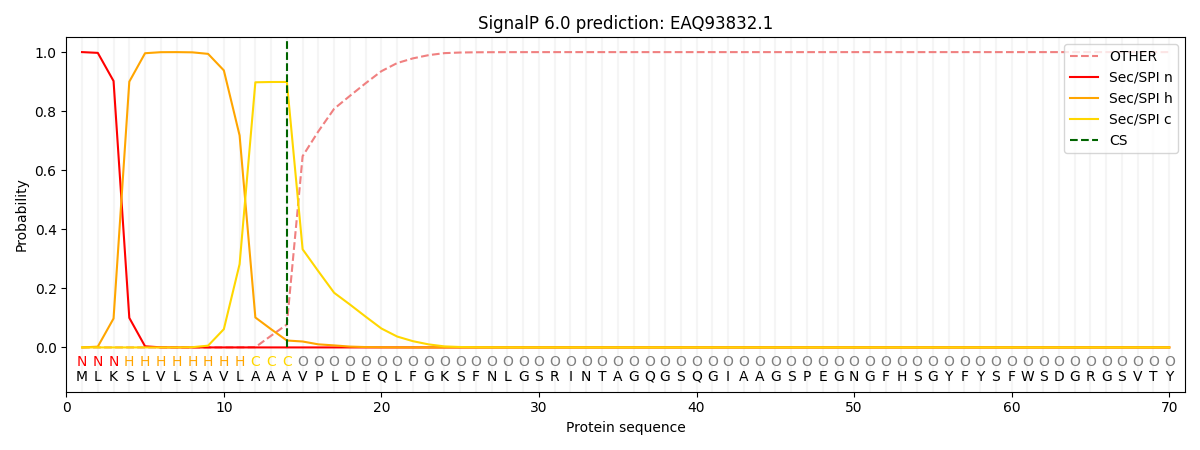
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000364 | 0.999610 | CS pos: 14-15. Pr: 0.8989 |
