You are browsing environment: FUNGIDB
CAZyme Information: CPSG_05530-t26_1-p1
You are here: Home > Sequence: CPSG_05530-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Coccidioides posadasii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Onygenaceae; Coccidioides; Coccidioides posadasii | |||||||||||
| CAZyme ID | CPSG_05530-t26_1-p1 | |||||||||||
| CAZy Family | GH3 | |||||||||||
| CAZyme Description | beta-glucosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH17 | 302 | 534 | 9.7e-18 | 0.9131832797427653 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 227625 | Scw11 | 8.48e-54 | 274 | 536 | 39 | 304 | Exo-beta-1,3-glucanase, GH17 family [Carbohydrate transport and metabolism]. |
| 275319 | predic_Ig_block | 1.52e-05 | 227 | 274 | 71 | 122 | putative immunoglobulin-blocking virulence protein. Members of this family are putative virulence proteins of Mycoplasma and Ureaplasma species. Members share a region of sequence similarity (see TIGR04524) with protein M, a Mycoplasma genitalium protein that binds a conserved light chain region of IgG and blocks its protective function of antigen-specific binding. The seed alignment for this model includes an N-terminal signal-anchor domain and a proline-rich linker domain, and a C-terminal extension, in addition to the protein M-like domain recognized by TIGR04524. [Cellular processes, Pathogenesis] |
| 411407 | PspC_subgroup_1 | 2.00e-05 | 227 | 293 | 420 | 502 | pneumococcal surface protein PspC, choline-binding form. The pneumococcal surface protein PspC, as described in Streptococcus pneumoniae, is a repetitive and highly variable protein, recognized by a conserved N-terminal domain and also by genomic location. This form, subgroup 1, has variable numbers of a choline-binding repeat in the C-terminal region, and is also known as choline-binding protein A. The other form, subgroup 2, is anchored covalently after cleavage by sortase at a C-terminal LPXTG site. |
| 223021 | PHA03247 | 3.29e-05 | 85 | 289 | 2776 | 2963 | large tegument protein UL36; Provisional |
| 411607 | Chlamy_inclu_1 | 6.78e-04 | 225 | 275 | 45 | 95 | inclusion-associated protein. Proteins of this family are inclusion-associated proteins in Chlamydia. It has been shown that protein CPj0783, which is identical to the HMM seed protein WP_010892266, was localized on Chlamydial inclusion. CPj0783 interacted with host Huntingtin-protein14, which may play an important role in disturbing the vesicle transport system to escape host lysosomal or autophagosomal degradation. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 537 | 1 | 537 | |
| 1.29e-151 | 33 | 536 | 104 | 592 | |
| 5.23e-145 | 1 | 536 | 1 | 601 | |
| 5.23e-145 | 1 | 536 | 1 | 601 | |
| 5.23e-145 | 1 | 536 | 1 | 601 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.29e-146 | 1 | 536 | 1 | 601 | Probable beta-glucosidase btgE OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=btgE PE=3 SV=1 |
|
| 9.29e-146 | 1 | 536 | 1 | 601 | Probable beta-glucosidase btgE OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=btgE PE=3 SV=1 |
|
| 3.88e-138 | 1 | 536 | 1 | 564 | Probable beta-glucosidase btgE OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=btgE PE=3 SV=2 |
|
| 3.88e-138 | 1 | 536 | 1 | 564 | Probable beta-glucosidase btgE OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=btgE PE=3 SV=2 |
|
| 1.06e-137 | 1 | 536 | 1 | 564 | Probable beta-glucosidase btgE OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=btgE PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
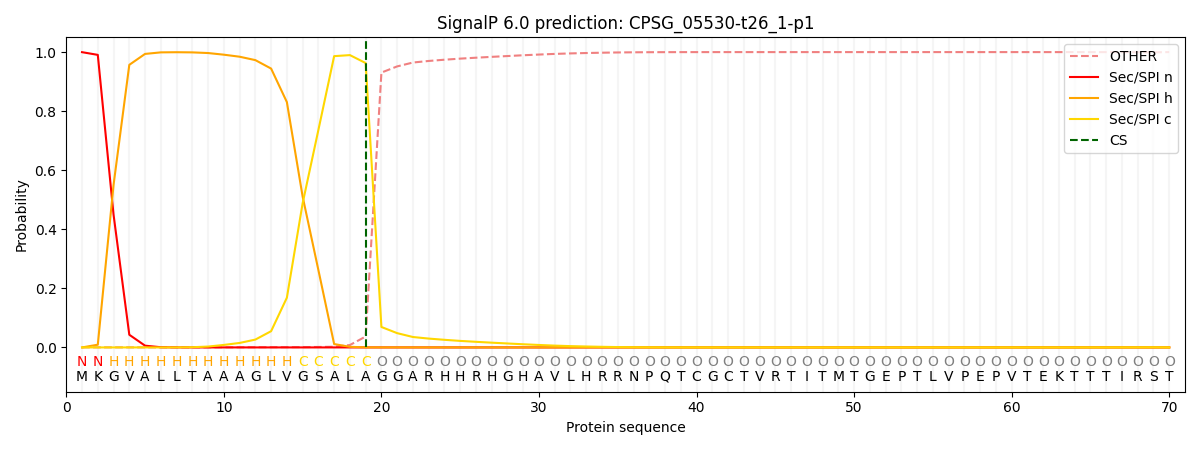
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.002673 | 0.997335 | CS pos: 19-20. Pr: 0.9629 |
