You are browsing environment: FUNGIDB
CAZyme Information: CLUG_05225-t34_1-p1
You are here: Home > Sequence: CLUG_05225-t34_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
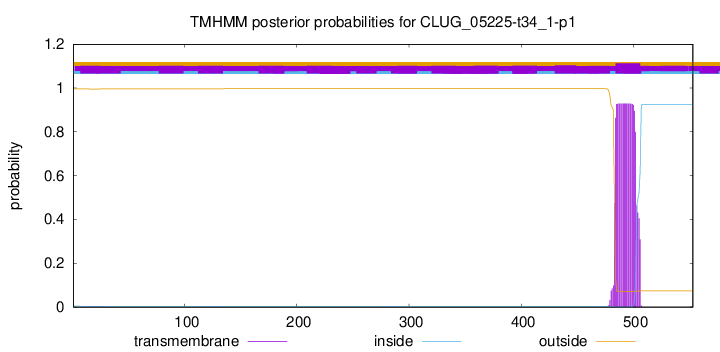
TMHMM annotations
Basic Information help
| Species | Clavispora lusitaniae | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Saccharomycetes; ; Metschnikowiaceae; Clavispora; Clavispora lusitaniae | |||||||||||
| CAZyme ID | CLUG_05225-t34_1-p1 | |||||||||||
| CAZy Family | GT66 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 1.10.3.2:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA1 | 13 | 298 | 1.3e-105 | 0.8011869436201781 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 259966 | CuRO_3_Fet3p | 3.39e-88 | 256 | 422 | 1 | 160 | The third Cupredoxin domain of multicopper oxidase Fet3p. Fet3p catalyzes the ferroxidase reaction, which couples the oxidation of Fe(II) to Fe(III) with the four-electron reduction of molecular oxygen to water. Fet3p is a type I membrane protein with the amino-terminal oxidase domain in the extracellular space and the carboxyl terminus in the cytoplasm. The periplasmic produced Fe(III) is transferred to the permease Ftr1p for import into the cytosol. The four copper ions are inserted post-translationally and are essential for catalytic activity, thus linking copper and iron homeostasis. Like other related multicopper oxidases (MCOs), Fet3p is composed of three cupredoxin domains that include one mononuclear and one trinuclear copper center. The copper ions are bound in several sites: Type 1, Type 2, and/or Type 3. The ensemble of types 2 and 3 copper is called a trinuclear cluster. MCOs oxidize their substrate by accepting electrons at a mononuclear copper center and transferring them to the active site trinuclear copper center. The cupredoxin domain 3 of 3-domain MCOs contains the Type 1 (T1) copper binding site and part the trinuclear copper binding site, which is located at the interface of domains 1 and 3. |
| 259945 | CuRO_2_Fet3p_like | 5.19e-67 | 67 | 209 | 5 | 146 | The second Cupredoxin domain of multicopper oxidase Fet3P. Fet3p catalyzes the ferroxidase reaction, which couples the oxidation of Fe(II) to Fe(III) with the four-electron reduction of molecular oxygen to water. Fet3p is a type I membrane protein with the amino-terminal oxidase domain in the extracellular space and the carboxyl terminus in the cytoplasm. The periplasmic produced Fe(III) is transferred to the permease Ftr1p for import into the cytosol. The four copper ions are inserted post-translationally and are essential for catalytic activity, thus linking copper and iron homeostasis. Like other related multicopper oxidases (MCOs), Fet3p is composed of three cupredoxin domains that include one mononuclear and one trinuclear copper center. The copper ions are bound in several sites: Type 1, Type 2, and/or Type 3. The ensemble of types 2 and 3 copper is called a trinuclear cluster. MCOs oxidize their substrate by accepting electrons at a mononuclear copper center and transferring them to the active site trinuclear copper center. The cupredoxin domain 2 of 3-domain MCOs has lost the ability to bind copper. |
| 225043 | SufI | 1.78e-63 | 18 | 422 | 117 | 450 | Multicopper oxidase with three cupredoxin domains (includes cell division protein FtsP and spore coat protein CotA) [Cell cycle control, cell division, chromosome partitioning, Inorganic ion transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
| 274555 | ascorbase | 1.44e-53 | 14 | 427 | 85 | 530 | L-ascorbate oxidase, plant type. Members of this protein family are the copper-containing enzyme L-ascorbate oxidase (EC 1.10.3.3), also called ascorbase. This family is found in flowering plants, and shows greater sequence similarity to a family of laccases (EC 1.10.3.2) from plants than to other known ascorbate oxidases. |
| 400194 | Cu-oxidase_2 | 1.02e-51 | 282 | 424 | 1 | 138 | Multicopper oxidase. This entry contains many divergent copper oxidase-like domains that are not recognized by the pfam00394 model. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 21 | 553 | 110 | 642 | |
| 0.0 | 21 | 553 | 110 | 642 | |
| 0.0 | 21 | 553 | 110 | 642 | |
| 0.0 | 21 | 553 | 110 | 642 | |
| 0.0 | 21 | 553 | 110 | 642 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.14e-157 | 15 | 478 | 88 | 534 | Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_B Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_C Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_D Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_E Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae],1ZPU_F Crystal Structure of Fet3p, a Multicopper Oxidase that Functions in Iron Import [Saccharomyces cerevisiae] |
|
| 4.95e-47 | 14 | 420 | 91 | 472 | Crystal structure of LacB from Trametes sp. AH28-2 [Trametes sp. AH28-2],3KW7_B Crystal structure of LacB from Trametes sp. AH28-2 [Trametes sp. AH28-2] |
|
| 1.14e-45 | 15 | 420 | 92 | 467 | CRYSTAL STRUCTURE OF A FULLY FUNCTIONAL LACCASE FROM THE LIGNINOLYTIC FUNGUS PYCNOPORUS CINNABARINUS [Trametes cinnabarina] |
|
| 1.52e-45 | 14 | 450 | 91 | 494 | Crystal Structure of Laccase from Cerrena sp. RSD1 [Cerrena],5Z1X_B Crystal Structure of Laccase from Cerrena sp. RSD1 [Cerrena],5Z22_A Crystal Structure of Laccase from Cerrena sp. RSD1 [Cerrena] |
|
| 5.51e-44 | 15 | 420 | 113 | 488 | Crystal structure of laccases from Pycnoporus sanguineus, izoform II [Trametes coccinea],5NQ9_A Crystal structure of laccases from Pycnoporus sanguineus, izoform II, monoclinic [Trametes sanguinea],5NQ9_C Crystal structure of laccases from Pycnoporus sanguineus, izoform II, monoclinic [Trametes sanguinea] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.20e-178 | 14 | 509 | 108 | 582 | Iron transport multicopper oxidase FET3 OS=Candida albicans OX=5476 GN=FET3 PE=3 SV=1 |
|
| 4.38e-172 | 14 | 515 | 112 | 597 | Iron transport multicopper oxidase FET3 OS=Kluyveromyces lactis (strain ATCC 8585 / CBS 2359 / DSM 70799 / NBRC 1267 / NRRL Y-1140 / WM37) OX=284590 GN=FET3 PE=3 SV=1 |
|
| 7.82e-166 | 15 | 548 | 109 | 626 | Iron transport multicopper oxidase FET3 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=FET3 PE=1 SV=2 |
|
| 6.07e-165 | 14 | 509 | 104 | 586 | Iron transport multicopper oxidase FET3 OS=Candida glabrata (strain ATCC 2001 / CBS 138 / JCM 3761 / NBRC 0622 / NRRL Y-65) OX=284593 GN=FET3 PE=3 SV=1 |
|
| 1.66e-161 | 15 | 527 | 111 | 614 | Iron transport multicopper oxidase FET5 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=FET5 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
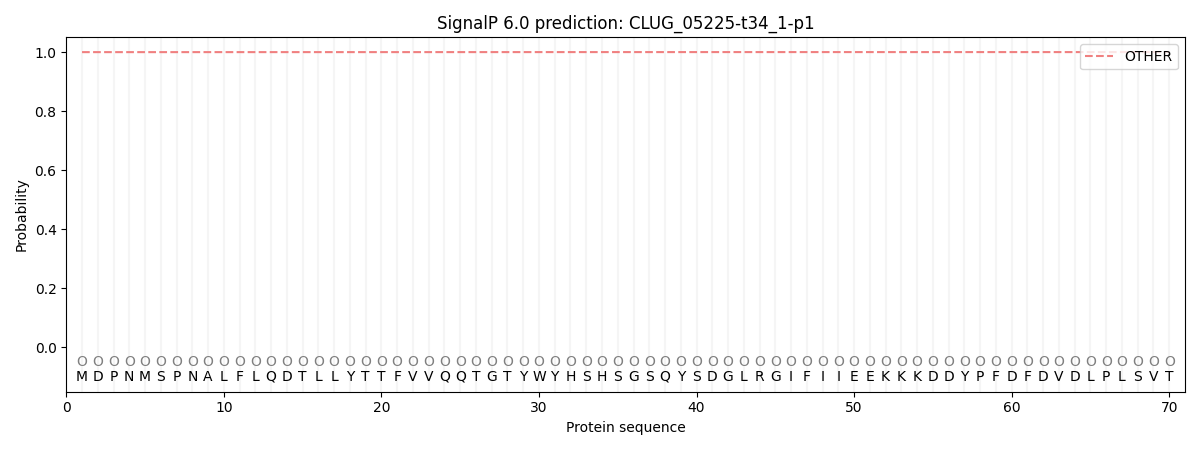
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999929 | 0.000111 |