You are browsing environment: FUNGIDB
CAZyme Information: CLCR_06677-t43_1-p1
You are here: Home > Sequence: CLCR_06677-t43_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Cladophialophora carrionii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Herpotrichiellaceae; Cladophialophora; Cladophialophora carrionii | |||||||||||
| CAZyme ID | CLCR_06677-t43_1-p1 | |||||||||||
| CAZy Family | GH5 | |||||||||||
| CAZyme Description | glycoside hydrolase family 31 protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.177:12 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH31 | 191 | 623 | 7.7e-67 | 0.9812646370023419 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 224418 | YicI | 8.67e-64 | 93 | 698 | 143 | 751 | Alpha-glucosidase, glycosyl hydrolase family GH31 [Carbohydrate transport and metabolism]. |
| 395838 | Glyco_hydro_31 | 3.05e-52 | 182 | 623 | 1 | 442 | Glycosyl hydrolases family 31. Glycosyl hydrolases are key enzymes of carbohydrate metabolism. Family 31 comprises of enzymes that are, or similar to, alpha- galactosidases. |
| 236731 | PRK10658 | 1.98e-34 | 145 | 620 | 198 | 665 | putative alpha-glucosidase; Provisional |
| 269878 | GH31_NET37 | 2.92e-34 | 268 | 576 | 47 | 360 | glucosidase NET37. NET37 (also known as KIAA1161) is a human lamina-associated nuclear envelope transmembrane protein. A member of the glycosyl hydrolase family 31 (GH31) , it has been shown to be required for myogenic differentiation of C2C12 cells. Related proteins are found in eukaryotes and prokaryotes. Enzymes of the GH31 family possess a wide range of different hydrolytic activities including alpha-glucosidase (glucoamylase and sucrase-isomaltase), alpha-xylosidase, 6-alpha-glucosyltransferase, 3-alpha-isomaltosyltransferase and alpha-1,4-glucan lyase. All GH31 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. |
| 236691 | PRK10426 | 2.20e-29 | 131 | 621 | 126 | 621 | alpha-glucosidase; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.36e-255 | 20 | 708 | 31 | 719 | |
| 2.36e-44 | 96 | 626 | 151 | 672 | |
| 4.47e-44 | 143 | 626 | 198 | 672 | |
| 5.24e-44 | 96 | 648 | 152 | 691 | |
| 6.01e-44 | 80 | 648 | 136 | 690 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.89e-31 | 68 | 711 | 125 | 740 | Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_B Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_C Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_D Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_E Structure of the YicI thiosugar Michaelis complex [Escherichia coli],2F2H_F Structure of the YicI thiosugar Michaelis complex [Escherichia coli] |
|
| 2.93e-31 | 68 | 711 | 125 | 740 | Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_B Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_C Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_D Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_E Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSI_F Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_A Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_B Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_C Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_D Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_E Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSJ_F Structure of a Family 31 alpha glycosidase [Escherichia coli],1XSK_A Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_B Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_C Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_D Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_E Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli],1XSK_F Structure of a Family 31 alpha glycosidase glycosyl-enzyme intermediate [Escherichia coli] |
|
| 2.38e-28 | 97 | 623 | 196 | 715 | Crystal structure of Family 31 alpha-glucosidase (BT_0339) from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482],5F7C_B Crystal structure of Family 31 alpha-glucosidase (BT_0339) from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482],5F7C_C Crystal structure of Family 31 alpha-glucosidase (BT_0339) from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482],5F7C_D Crystal structure of Family 31 alpha-glucosidase (BT_0339) from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482] |
|
| 4.25e-25 | 94 | 650 | 136 | 705 | Crystal structure of Paenibacillus sp. 598K alpha-1,6-glucosyltransferase [Paenibacillus sp. 598K],5X7O_B Crystal structure of Paenibacillus sp. 598K alpha-1,6-glucosyltransferase [Paenibacillus sp. 598K],5X7P_A Crystal structure of Paenibacillus sp. 598K alpha-1,6-glucosyltransferase complexed with acarbose [Paenibacillus sp. 598K],5X7P_B Crystal structure of Paenibacillus sp. 598K alpha-1,6-glucosyltransferase complexed with acarbose [Paenibacillus sp. 598K],5X7Q_A Crystal structure of Paenibacillus sp. 598K alpha-1,6-glucosyltransferase complexed with maltohexaose [Paenibacillus sp. 598K],5X7Q_B Crystal structure of Paenibacillus sp. 598K alpha-1,6-glucosyltransferase complexed with maltohexaose [Paenibacillus sp. 598K],5X7R_A Crystal structure of Paenibacillus sp. 598K alpha-1,6-glucosyltransferase complexed with isomaltohexaose [Paenibacillus sp. 598K],5X7R_B Crystal structure of Paenibacillus sp. 598K alpha-1,6-glucosyltransferase complexed with isomaltohexaose [Paenibacillus sp. 598K],5X7S_A Crystal structure of Paenibacillus sp. 598K alpha-1,6-glucosyltransferase, terbium derivative [Paenibacillus sp. 598K],5X7S_B Crystal structure of Paenibacillus sp. 598K alpha-1,6-glucosyltransferase, terbium derivative [Paenibacillus sp. 598K] |
|
| 1.04e-23 | 143 | 682 | 289 | 865 | Cycloalternan-forming enzyme from Listeria monocytogenes in complex with pentasaccharide substrate [Listeria monocytogenes EGD-e] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 7.06e-33 | 92 | 622 | 167 | 695 | Alpha-xylosidase OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=agdD PE=1 SV=1 |
|
| 2.64e-31 | 38 | 627 | 92 | 678 | Alpha-xylosidase XylQ OS=Lactiplantibacillus pentosus OX=1589 GN=xylQ PE=1 SV=1 |
|
| 1.48e-30 | 68 | 711 | 125 | 740 | Alpha-xylosidase OS=Escherichia coli (strain K12) OX=83333 GN=yicI PE=1 SV=2 |
|
| 2.55e-21 | 144 | 628 | 193 | 679 | Alpha-glucosidase 2 OS=Bacillus thermoamyloliquefaciens OX=1425 PE=3 SV=1 |
|
| 1.91e-19 | 402 | 623 | 601 | 810 | Neutral alpha-glucosidase AB OS=Homo sapiens OX=9606 GN=GANAB PE=1 SV=3 |
SignalP and Lipop Annotations help
This protein is predicted as SP
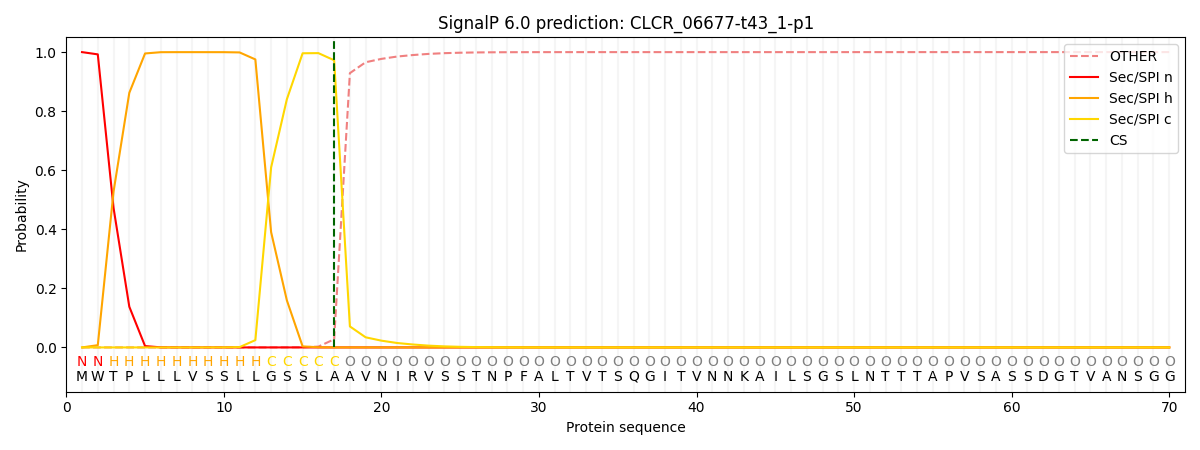
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000348 | 0.999608 | CS pos: 17-18. Pr: 0.9724 |
