You are browsing environment: FUNGIDB
CAZyme Information: CLCR_03411-t43_1-p1
You are here: Home > Sequence: CLCR_03411-t43_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Cladophialophora carrionii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Herpotrichiellaceae; Cladophialophora; Cladophialophora carrionii | |||||||||||
| CAZyme ID | CLCR_03411-t43_1-p1 | |||||||||||
| CAZy Family | GH17 | |||||||||||
| CAZyme Description | Alpha | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.15:96 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT20 | 15 | 502 | 1.5e-163 | 0.9705263157894737 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 340820 | GT20_TPS | 0.0 | 17 | 502 | 1 | 462 | trehalose-6-phosphate synthase. Trehalose-6-Phosphate Synthase (TPS, EC 2.4.1.15) is a glycosyltransferase that catalyses the synthesis of alpha,alpha-1,1-trehalose-6-phosphate from glucose-6-phosphate using a UDP-glucose donor. It is a key enzyme in the trehalose synthesis pathway. Trehalose is a nonreducing disaccharide present in a wide variety of organisms and may serve as a source of energy and carbon. It is characterized most notably in insect, plant, and microbial cells. Its production is often associated with a variety of stress conditions, including desiccation, dehydration, heat, cold, and oxidation. This family represents the catalytic domain of the TPS. Some members of this domain family coexist with a C-terminal trehalose phosphatase domain. |
| 395781 | Glyco_transf_20 | 0.0 | 16 | 500 | 1 | 466 | Glycosyltransferase family 20. Members of this family belong to glycosyl transferase family 20. OtsA (Trehalose-6-phosphate synthase) is homologous to regions in the subunits of yeast trehalose-6-phosphate synthase/phosphate complex,. |
| 274112 | trehalose_OtsA | 6.23e-176 | 17 | 503 | 1 | 455 | alpha,alpha-trehalose-phosphate synthase [UDP-forming]. This enzyme catalyzes the key, penultimate step in biosynthesis of trehalose, a compatible solute made as an osmoprotectant in some species in all three domains of life. The gene symbol OtsA stands for osmotically regulated trehalose synthesis A. Trehalose helps protect against both osmotic and thermal stresses, and is made from two glucose subunits. This model excludes glucosylglycerol-phosphate synthase, an enzyme of an analogous osmoprotectant system in many cyanobacterial strains. This model does not identify archaeal examples, as they are more divergent than glucosylglycerol-phosphate synthase. Sequences that score in the gray zone between the trusted and noise cutoffs include a number of yeast multidomain proteins in which the N-terminal domain may be functionally equivalent to this family. The gray zone also includes the OtsA of Cornyebacterium glutamicum (and related species), shown to be responsible for synthesis of only trace amounts of trehalose while the majority is synthesized by the TreYZ pathway; the significance of OtsA in this species is unclear (see Wolf, et al., ). [Cellular processes, Adaptations to atypical conditions] |
| 223457 | OtsA | 2.52e-167 | 16 | 504 | 15 | 479 | Trehalose-6-phosphate synthase [Carbohydrate transport and metabolism]. |
| 184712 | PRK14501 | 2.09e-157 | 16 | 509 | 1 | 467 | putative bifunctional trehalose-6-phosphate synthase/HAD hydrolase subfamily IIB; Provisional |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.88e-168 | 16 | 509 | 13 | 484 | |
| 5.50e-168 | 16 | 509 | 13 | 484 | |
| 1.04e-167 | 8 | 513 | 9 | 493 | |
| 1.11e-167 | 4 | 513 | 5 | 494 | |
| 3.24e-167 | 16 | 509 | 76 | 547 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.71e-170 | 15 | 511 | 3 | 476 | Structure of Candida albicans trehalose-6-phosphate synthase E341R/E346R in complex with UDP-glucose [Candida albicans SC5314],5HUV_B Structure of Candida albicans trehalose-6-phosphate synthase E341R/E346R in complex with UDP-glucose [Candida albicans SC5314] |
|
| 1.21e-168 | 15 | 511 | 3 | 476 | Structure of Candida albicans trehalose-6-phosphate synthase in complex with UDP-glucose [Candida albicans SC5314],5HUT_B Structure of Candida albicans trehalose-6-phosphate synthase in complex with UDP-glucose [Candida albicans SC5314],5HUU_A Structure of Candida albicans trehalose-6-phosphate synthase in complex with UDP and glucose-6-phosphate [Candida albicans SC5314],5HUU_B Structure of Candida albicans trehalose-6-phosphate synthase in complex with UDP and glucose-6-phosphate [Candida albicans SC5314],5HVL_A Structure of Candida albicans trehalose-6-phosphate synthase in complex with UDP and validoxylamine A [Candida albicans SC5314],5HVL_B Structure of Candida albicans trehalose-6-phosphate synthase in complex with UDP and validoxylamine A [Candida albicans SC5314] |
|
| 1.07e-160 | 1 | 499 | 1 | 471 | Structure of Aspergillus fumigatus trehalose-6-phosphate synthase A in complex with UDP and validoxylamine A [Aspergillus fumigatus Af293],5HVM_B Structure of Aspergillus fumigatus trehalose-6-phosphate synthase A in complex with UDP and validoxylamine A [Aspergillus fumigatus Af293] |
|
| 5.20e-160 | 13 | 503 | 8 | 475 | Structure of Aspergillus fumigatus trehalose-6-phosphate synthase B in complex with UDP and validoxylamine A [Aspergillus fumigatus Af293],5HVO_B Structure of Aspergillus fumigatus trehalose-6-phosphate synthase B in complex with UDP and validoxylamine A [Aspergillus fumigatus Af293],5HVO_C Structure of Aspergillus fumigatus trehalose-6-phosphate synthase B in complex with UDP and validoxylamine A [Aspergillus fumigatus Af293],5HVO_D Structure of Aspergillus fumigatus trehalose-6-phosphate synthase B in complex with UDP and validoxylamine A [Aspergillus fumigatus Af293] |
|
| 1.11e-155 | 17 | 499 | 1 | 459 | Structure of Tps1 apo structure [Pyricularia oryzae 70-15],6JBI_B Structure of Tps1 apo structure [Pyricularia oryzae 70-15],6JBR_A Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_B Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_D Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_F Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_H Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_K Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_M Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBR_O Tps1/UDP/T6P complex [Pyricularia oryzae 70-15],6JBW_A Structure of Tps1/UDP complex [Pyricularia oryzae 70-15],6JBW_B Structure of Tps1/UDP complex [Pyricularia oryzae 70-15] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.22e-168 | 15 | 511 | 3 | 476 | Alpha,alpha-trehalose-phosphate synthase [UDP-forming] OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=TPS1 PE=1 SV=1 |
|
| 2.63e-164 | 13 | 502 | 13 | 480 | Alpha,alpha-trehalose-phosphate synthase [UDP-forming] 56 kDa subunit OS=Kluyveromyces lactis (strain ATCC 8585 / CBS 2359 / DSM 70799 / NBRC 1267 / NRRL Y-1140 / WM37) OX=284590 GN=TPS1 PE=1 SV=1 |
|
| 1.24e-161 | 16 | 502 | 16 | 480 | Alpha,alpha-trehalose-phosphate synthase [UDP-forming] 56 kDa subunit OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=TPS1 PE=1 SV=2 |
|
| 4.75e-161 | 15 | 499 | 11 | 471 | Alpha,alpha-trehalose-phosphate synthase [UDP-forming] OS=Emericella nidulans (strain FGSC A4 / ATCC 38163 / CBS 112.46 / NRRL 194 / M139) OX=227321 GN=tpsA PE=3 SV=1 |
|
| 5.53e-160 | 1 | 499 | 1 | 471 | Alpha,alpha-trehalose-phosphate synthase [UDP-forming] 1 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=tpsA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
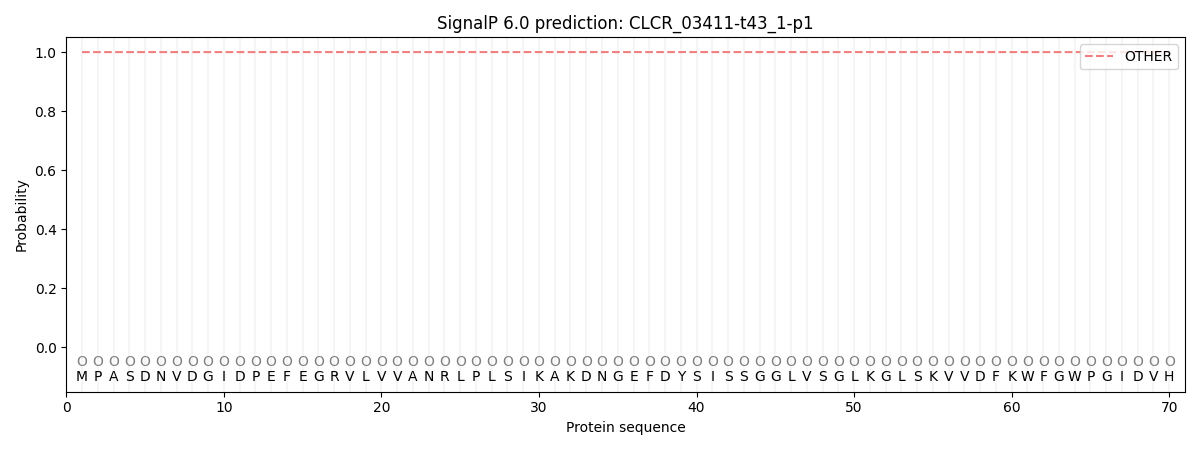
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000055 | 0.000000 |
