You are browsing environment: FUNGIDB
CAZyme Information: CLCR_03001-t43_1-p1
You are here: Home > Sequence: CLCR_03001-t43_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Cladophialophora carrionii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Herpotrichiellaceae; Cladophialophora; Cladophialophora carrionii | |||||||||||
| CAZyme ID | CLCR_03001-t43_1-p1 | |||||||||||
| CAZy Family | GH154 | |||||||||||
| CAZyme Description | Initiation-specific alpha-1,6-mannosyltransferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.257:12 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT32 | 115 | 193 | 2.9e-20 | 0.8888888888888888 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 226297 | OCH1 | 8.50e-43 | 68 | 355 | 61 | 345 | Mannosyltransferase OCH1 or related enzyme [Cell wall/membrane/envelope biogenesis]. |
| 398274 | Gly_transf_sug | 8.27e-19 | 116 | 193 | 10 | 88 | Glycosyltransferase sugar-binding region containing DXD motif. The DXD motif is a short conserved motif found in many families of glycosyltransferases, which add a range of different sugars to other sugars, phosphates and proteins. DXD-containing glycosyltransferases all use nucleoside diphosphate sugars as donors and require divalent cations, usually manganese. The DXD motif is expected to play a carbohydrate binding role in sugar-nucleoside diphosphate and manganese dependent glycosyltransferases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.58e-151 | 6 | 360 | 3 | 372 | |
| 1.58e-151 | 6 | 360 | 3 | 372 | |
| 5.57e-151 | 51 | 360 | 49 | 356 | |
| 6.87e-151 | 6 | 355 | 3 | 354 | |
| 3.68e-150 | 6 | 355 | 3 | 353 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.97e-95 | 72 | 355 | 82 | 376 | Initiation-specific alpha-1,6-mannosyltransferase OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=OCH1 PE=3 SV=1 |
|
| 3.48e-85 | 59 | 355 | 120 | 394 | Initiation-specific alpha-1,6-mannosyltransferase OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=och1 PE=1 SV=2 |
|
| 3.84e-77 | 72 | 360 | 79 | 475 | Initiation-specific alpha-1,6-mannosyltransferase OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=OCH1 PE=1 SV=1 |
|
| 2.62e-72 | 62 | 355 | 103 | 388 | Putative glycosyltransferase HOC1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=HOC1 PE=1 SV=3 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
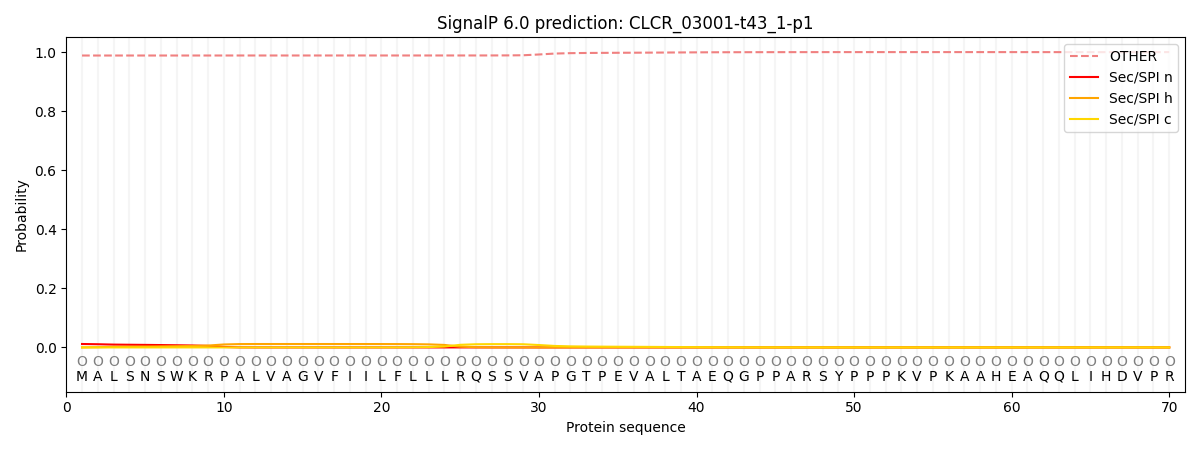
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.989077 | 0.010948 |
