You are browsing environment: FUNGIDB
CAZyme Information: CIMG_03293-t26_1-p1
You are here: Home > Sequence: CIMG_03293-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Coccidioides immitis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Onygenaceae; Coccidioides; Coccidioides immitis | |||||||||||
| CAZyme ID | CIMG_03293-t26_1-p1 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | sugar 1,4-lactone oxidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA7 | 31 | 239 | 3.6e-30 | 0.4606986899563319 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 273751 | FAD_lactone_ox | 1.10e-143 | 32 | 476 | 7 | 436 | sugar 1,4-lactone oxidases. This model represents a family of at least two different sugar 1,4 lactone oxidases, both involved in synthesizing ascorbic acid or a derivative. These include L-gulonolactone oxidase (EC 1.1.3.8) from rat and D-arabinono-1,4-lactone oxidase (EC 1.1.3.37) from Saccharomyces cerevisiae. Members are proposed to have the cofactor FAD covalently bound at a site specified by Prosite motif PS00862; OX2_COVAL_FAD; 1. |
| 397924 | ALO | 2.59e-108 | 199 | 482 | 1 | 258 | D-arabinono-1,4-lactone oxidase. This domain is specific to D-arabinono-1,4-lactone oxidase EC:1.1.3.-, which is involved in the final step of the D-erythroascorbic acid biosynthesis pathway. |
| 223354 | GlcD | 5.31e-44 | 30 | 475 | 21 | 450 | FAD/FMN-containing dehydrogenase [Energy production and conversion]. |
| 215258 | PLN02465 | 5.80e-35 | 32 | 479 | 89 | 565 | L-galactono-1,4-lactone dehydrogenase |
| 396238 | FAD_binding_4 | 2.94e-34 | 40 | 175 | 1 | 139 | FAD binding domain. This family consists of various enzymes that use FAD as a co-factor, most of the enzymes are similar to oxygen oxidoreductase. One of the enzymes Vanillyl-alcohol oxidase (VAO) has a solved structure, the alignment includes the FAD binding site, called the PP-loop, between residues 99-110. The FAD molecule is covalently bound in the known structure, however the residue that links to the FAD is not in the alignment. VAO catalyzes the oxidation of a wide variety of substrates, ranging form aromatic amines to 4-alkylphenols. Other members of this family include D-lactate dehydrogenase, this enzyme catalyzes the conversion of D-lactate to pyruvate using FAD as a co-factor; mitomycin radical oxidase, this enzyme oxidizes the reduced form of mitomycins and is involved in mitomycin resistance. This family includes MurB an UDP-N-acetylenolpyruvoylglucosamine reductase enzyme EC:1.1.1.158. This enzyme is involved in the biosynthesis of peptidoglycan. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.36e-09 | 40 | 236 | 59 | 267 | |
| 1.68e-09 | 16 | 202 | 107 | 300 | |
| 3.38e-09 | 40 | 198 | 62 | 223 | |
| 4.24e-09 | 39 | 205 | 212 | 381 | |
| 4.35e-09 | 39 | 222 | 48 | 239 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.86e-28 | 27 | 249 | 110 | 331 | Assembly intermediate of the plant mitochondrial complex I [Brassica oleracea] |
|
| 3.40e-15 | 32 | 259 | 13 | 238 | Alditol Oxidase from Streptomyces coelicolor A3(2): Native Enzyme [Streptomyces coelicolor A3(2)],2VFS_A Alditol Oxidase from Streptomyces coelicolor A3(2): Complex with Xylitol [Streptomyces coelicolor A3(2)],2VFT_A Alditol Oxidase from Streptomyces coelicolor A3(2): Complex with Sorbitol [Streptomyces coelicolor A3(2)],2VFU_A Alditol Oxidase from Streptomyces coelicolor A3(2): Complex with Mannitol [Streptomyces coelicolor A3(2)],2VFV_A Alditol Oxidase from Streptomyces coelicolor A3(2): Complex with Sulphite [Streptomyces coelicolor A3(2)] |
|
| 8.45e-11 | 40 | 236 | 37 | 245 | Crystal structure of carbohydrate oxidase from Microdochium nivale [Microdochium nivale],3RJA_A Crystal structure of carbohydrate oxidase from Microdochium nivale in complex with substrate analogue [Microdochium nivale] |
|
| 1.99e-10 | 40 | 200 | 48 | 212 | Physcomitrella patens BBE-like 1 wild-type [Physcomitrium patens],6EO4_B Physcomitrella patens BBE-like 1 wild-type [Physcomitrium patens] |
|
| 1.99e-10 | 40 | 200 | 48 | 212 | Physcomitrella patens BBE-like 1 variant D396N [Physcomitrium patens],6EO5_B Physcomitrella patens BBE-like 1 variant D396N [Physcomitrium patens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.24e-203 | 13 | 530 | 24 | 556 | Putative D-arabinono-1,4-lactone oxidase OS=Neurospora crassa (strain ATCC 24698 / 74-OR23-1A / CBS 708.71 / DSM 1257 / FGSC 987) OX=367110 GN=alo-1 PE=3 SV=1 |
|
| 1.88e-126 | 23 | 476 | 9 | 514 | D-arabinono-1,4-lactone oxidase OS=Yarrowia lipolytica (strain CLIB 122 / E 150) OX=284591 GN=ALO1 PE=3 SV=1 |
|
| 7.28e-109 | 20 | 476 | 10 | 543 | D-arabinono-1,4-lactone oxidase OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=ALO1 PE=1 SV=1 |
|
| 6.05e-107 | 27 | 476 | 16 | 513 | D-arabinono-1,4-lactone oxidase OS=Ashbya gossypii (strain ATCC 10895 / CBS 109.51 / FGSC 9923 / NRRL Y-1056) OX=284811 GN=ALO1 PE=3 SV=1 |
|
| 3.86e-106 | 32 | 479 | 19 | 516 | D-arabinono-1,4-lactone oxidase OS=Kluyveromyces lactis (strain ATCC 8585 / CBS 2359 / DSM 70799 / NBRC 1267 / NRRL Y-1140 / WM37) OX=284590 GN=ALO1 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
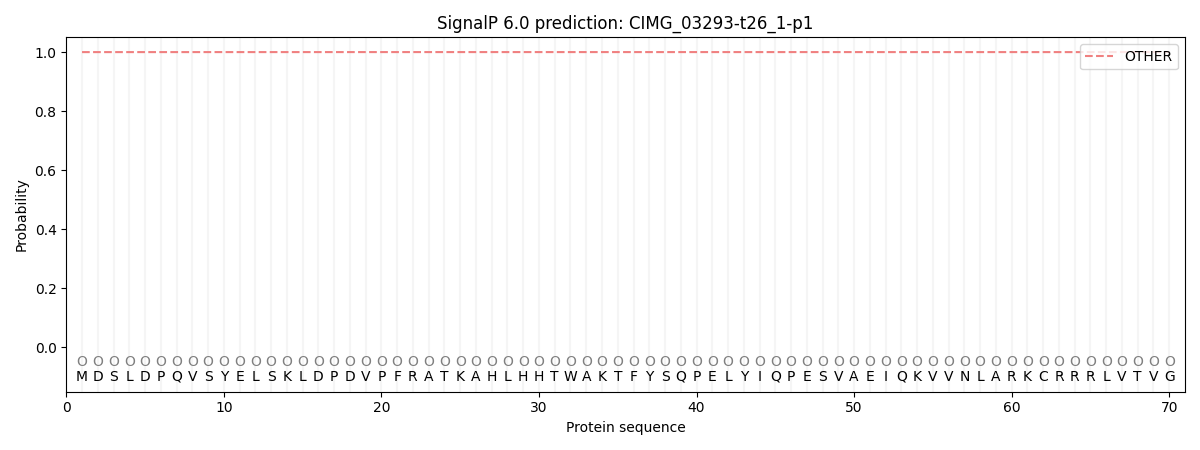
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000039 | 0.000000 |
