You are browsing environment: FUNGIDB
CAZyme Information: CGB_J2210C-t26_1-p1
You are here: Home > Sequence: CGB_J2210C-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
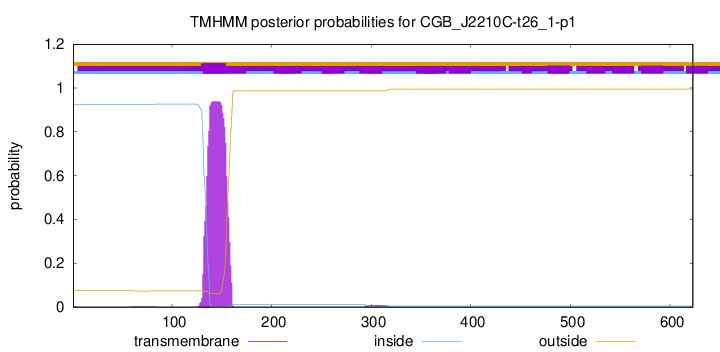
TMHMM annotations
Basic Information help
| Species | Cryptococcus gattii VGI | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Arthropoda; Insecta; ; Eriococcidae; Cryptococcus; Cryptococcus gattii VGI | |||||||||||
| CAZyme ID | CGB_J2210C-t26_1-p1 | |||||||||||
| CAZy Family | GT32 | |||||||||||
| CAZyme Description | putative glucosidase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH16 | 218 | 571 | 1.9e-137 | 0.9969325153374233 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 397841 | SKN1 | 0.0 | 116 | 623 | 24 | 500 | Beta-glucan synthesis-associated protein (SKN1). This family consists of the beta-glucan synthesis-associated proteins KRE6 and SKN1. Beta1,6-Glucan is a key component of the yeast cell wall, interconnecting cell wall proteins, beta1,3-glucan, and chitin. It has been postulated that the synthesis of beta1,6-glucan begins in the endoplasmic reticulum with the formation of protein-bound primer structures and that these primer structures are extended in the Golgi complex by two putative glucosyltransferases that are functionally redundant, Kre6 and Skn1. This is followed by maturation steps at the cell surface and by coupling to other cell wall macromolecules. |
| 185689 | GH16_fungal_KRE6_glucanase | 2.12e-167 | 214 | 572 | 1 | 295 | Saccharomyces cerevisiae KRE6 and related glucanses, member of glycosyl hydrolase family 16. KRE6 is a Saccharomyces cerevisiae glucanase that participates in the synthesis of beta-1,6-glucan, a major structural component of the cell wall. It is a golgi membrane protein required for normal beta-1,6-glucan levels in the cell wall. KRE6 is closely realted to laminarinase, a glycosyl hydrolase family 16 member that hydrolyzes 1,3-beta-D-glucosidic linkages in 1,3-beta-D-glucans such as laminarins, curdlans, paramylons, and pachymans, with very limited action on mixed-link (1,3-1,4-)-beta-D-glucans. |
| 225182 | BglS | 2.53e-42 | 192 | 585 | 29 | 355 | Beta-glucanase, GH16 family [Carbohydrate transport and metabolism]. |
| 185693 | GH16_laminarinase_like | 1.68e-18 | 216 | 571 | 1 | 235 | Laminarinase, member of the glycosyl hydrolase family 16. Laminarinase, also known as glucan endo-1,3-beta-D-glucosidase, is a glycosyl hydrolase family 16 member that hydrolyzes 1,3-beta-D-glucosidic linkages in 1,3-beta-D-glucans such as laminarins, curdlans, paramylons, and pachymans, with very limited action on mixed-link (1,3-1,4-)-beta-D-glucans. |
| 185687 | GH16_beta_agarase | 1.91e-04 | 208 | 312 | 15 | 117 | Beta-agarase, member of glycosyl hydrolase family 16. Beta-agarase is a glycosyl hydrolase family 16 (GH16) member that hydrolyzes the internal beta-1,4-linkage of agarose, a hydrophilic polysaccharide found in the cell wall of Rhodophyceaea, marine red algae. Agarose is a linear chain of galactose units linked by alternating L-alpha-1,3- and D-beta-1,4-linkages that are additionally modified by a 3,6-anhydro-bridge. Agarose forms thermo-reversible gels that are widely used in the food industry or as a laboratory medium. While beta-agarases are also found in two other families derived from the sequence-based classification of glycosyl hydrolases (GH50, and GH86) the GH16 members are most abundant. This domain adopts a curved beta-sandwich conformation, with a tunnel-shaped active site cavity, referred to as a jellyroll fold. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 623 | 1 | 623 | |
| 0.0 | 1 | 623 | 1 | 623 | |
| 0.0 | 1 | 623 | 1 | 622 | |
| 0.0 | 1 | 623 | 1 | 622 | |
| 0.0 | 1 | 623 | 1 | 622 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.00e-134 | 146 | 621 | 293 | 736 | Beta-glucan synthesis-associated protein SKN1 OS=Candida albicans OX=5476 GN=SKN1 PE=2 SV=1 |
|
| 9.77e-134 | 101 | 623 | 214 | 711 | Beta-glucan synthesis-associated protein KRE6 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=KRE6 PE=1 SV=2 |
|
| 5.58e-133 | 106 | 623 | 259 | 763 | Beta-glucan synthesis-associated protein SKN1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=SKN1 PE=1 SV=1 |
|
| 8.93e-130 | 183 | 621 | 321 | 739 | Beta-glucan synthesis-associated protein KRE6 OS=Candida albicans OX=5476 GN=KRE6 PE=2 SV=1 |
|
| 5.07e-125 | 192 | 623 | 218 | 627 | Uncharacterized beta-glucan synthesis-associated protein C23H3.11c OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPAC23H3.11c PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
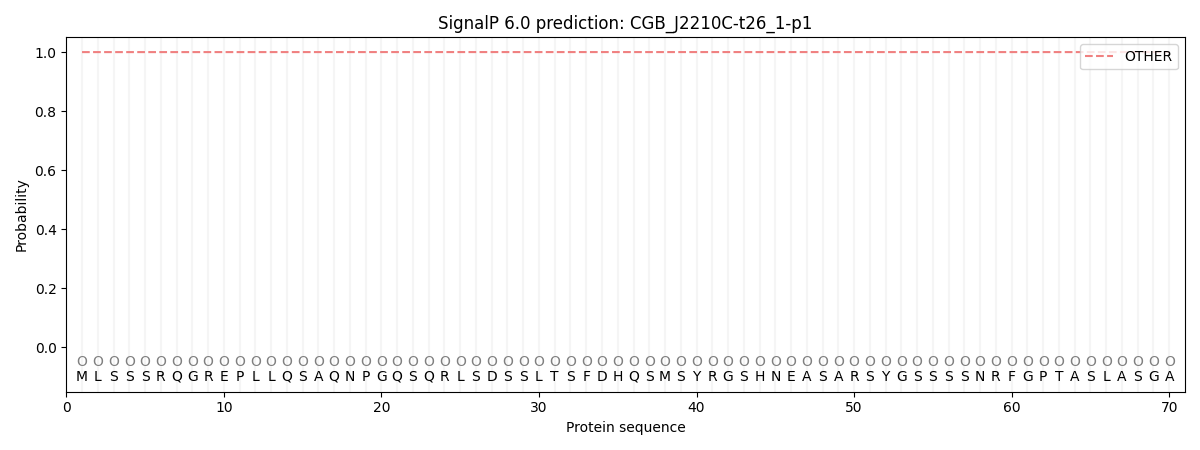
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000060 | 0.000000 |