You are browsing environment: FUNGIDB
CAZyme Information: CDH49878.1
You are here: Home > Sequence: CDH49878.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
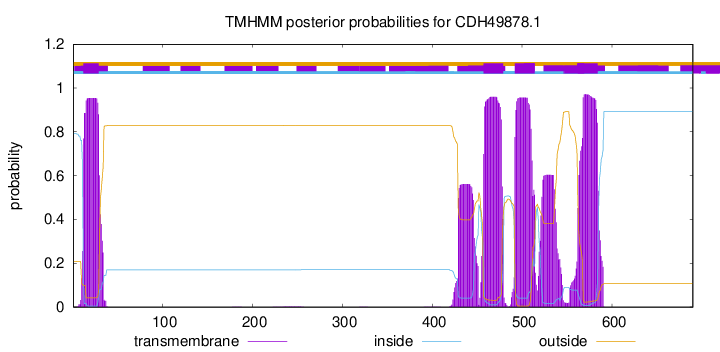
TMHMM annotations
Basic Information help
| Species | Lichtheimia corymbifera | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Mucoromycota; Mucoromycetes; ; Lichtheimiaceae; Lichtheimia; Lichtheimia corymbifera | |||||||||||
| CAZyme ID | CDH49878.1 | |||||||||||
| CAZy Family | CE4 | |||||||||||
| CAZyme Description | glycosyltransferase family 15 protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 2.4.1.131:20 | 2.4.1.-:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT15 | 73 | 343 | 2.1e-82 | 0.978021978021978 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 227353 | KTR1 | 7.19e-101 | 67 | 378 | 76 | 384 | Mannosyltransferase [Carbohydrate transport and metabolism]. |
| 396385 | Glyco_transf_15 | 2.69e-87 | 56 | 342 | 27 | 309 | Glycolipid 2-alpha-mannosyltransferase. This is a family of alpha-1,2 mannosyl-transferases involved in N-linked and O-linked glycosylation of proteins. Some of the enzymes in this family have been shown to be involved in O- and N-linked glycan modifications in the Golgi. |
| 270465 | CUE | 2.85e-07 | 649 | 686 | 1 | 38 | CUE domain found in ubiquitin-binding CUE proteins. This family includes many coupling of ubiquitin conjugation to endoplasmic reticulum degradation (CUE) domain containing proteins that are characterized by an FP and a di-leucine-like sequence and bind to monoubiquitin with varying affinities. Some higher eukaryotic CUE domain proteins do not bind monoubiquitin efficiently, since they carry LP, rather than FP among CUE domains. CUE domains form three-helix bundle structures and are distantly related to the ubiquitin-associated (UBA) domains which are widely occurring ubiquitin-binding motifs found in a broad range of cellular proteins in species ranging from yeast to human. The majority of family members contain one CUE domain, but some family members from fungi harbor two CUE domains. |
| 397127 | CUE | 6.36e-05 | 649 | 688 | 2 | 41 | CUE domain. CUE domains have been shown to bind ubiquitin. It has been suggested that CUE domains are related to pfam00627 and this has been confirmed by the structure of the domain. CUE domains also occur in two protein of the IL-1 signal transduction pathway, tollip and TAB2. |
| 270557 | CUE1_Cue2p_like | 1.82e-04 | 649 | 687 | 1 | 39 | CUE1 domain found in yeast ubiquitin-binding protein CUE2 (Cue2p) and similar proteins. Cue2p, also called coupling of ubiquitin conjugation to ER degradation protein 2, is encoded by the open reading frame (ORF) YKL090W. It is involved in the intramolecular monoubiquitination that serves as a regulatory signal in a variety of cellular processes in yeast. Cue2p contains two tandem CUE domains at the N-terminus. Both of them can bind monoubiquitin independently. This model corresponds to the first CUE domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.70e-260 | 1 | 378 | 1 | 371 | |
| 8.41e-151 | 61 | 377 | 55 | 369 | |
| 3.67e-83 | 73 | 377 | 69 | 368 | |
| 5.09e-77 | 65 | 376 | 76 | 386 | |
| 5.09e-77 | 65 | 376 | 76 | 386 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.40e-67 | 74 | 376 | 29 | 327 | Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p [Saccharomyces cerevisiae],1S4N_B Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p [Saccharomyces cerevisiae],1S4O_A Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p: binary complex with GDP/Mn [Saccharomyces cerevisiae],1S4O_B Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p: binary complex with GDP/Mn [Saccharomyces cerevisiae],1S4P_A Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p: ternary complex with GDP/Mn and methyl-alpha-mannoside acceptor [Saccharomyces cerevisiae],1S4P_B Crystal structure of yeast alpha1,2-mannosyltransferase Kre2p/Mnt1p: ternary complex with GDP/Mn and methyl-alpha-mannoside acceptor [Saccharomyces cerevisiae] |
|
| 9.67e-50 | 73 | 367 | 68 | 362 | Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_B Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_C Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_D Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_E Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOO_F Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, apo form [Aspergillus fumigatus A1163],7BOP_A Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_B Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_C Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_D Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_E Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163],7BOP_F Crystal Structure of Core-mannan synthase A (CmsA/Ktr4) from Aspergillus fumigatus, Mn/GDP-form [Aspergillus fumigatus A1163] |
|
| 3.74e-44 | 87 | 362 | 116 | 398 | X-ray structure of the mannosyltransferase Ktr4p from S. cerevisiae in complex with GDP [Saccharomyces cerevisiae S288C],5A07_B X-ray structure of the mannosyltransferase Ktr4p from S. cerevisiae in complex with GDP [Saccharomyces cerevisiae S288C],5A08_A X-ray structure of the mannosyltransferase Ktr4p from S. cerevisiae [Saccharomyces cerevisiae S288C],5A08_B X-ray structure of the mannosyltransferase Ktr4p from S. cerevisiae [Saccharomyces cerevisiae S288C] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.70e-73 | 74 | 376 | 142 | 441 | Glycolipid 2-alpha-mannosyltransferase 2 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=MNT2 PE=3 SV=4 |
|
| 5.87e-69 | 68 | 377 | 66 | 370 | O-glycoside alpha-1,2-mannosyltransferase omh1 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=omh1 PE=3 SV=1 |
|
| 9.48e-68 | 69 | 376 | 106 | 410 | Glycolipid 2-alpha-mannosyltransferase 1 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=MNT1 PE=3 SV=1 |
|
| 9.43e-66 | 74 | 376 | 123 | 421 | Glycolipid 2-alpha-mannosyltransferase OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=KRE2 PE=1 SV=1 |
|
| 9.66e-66 | 58 | 376 | 57 | 372 | Alpha-1,2 mannosyltransferase KTR1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=KTR1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
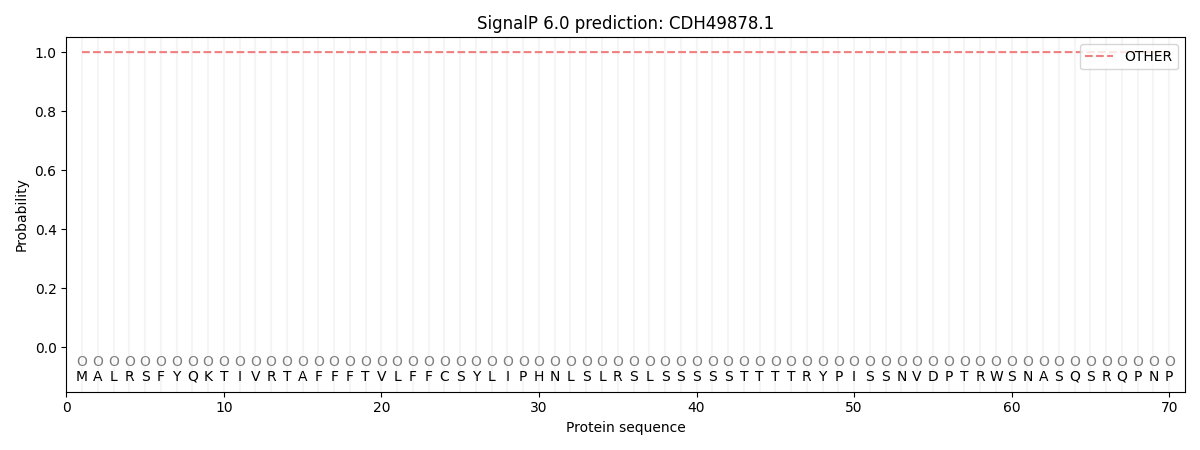
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999943 | 0.000053 |