You are browsing environment: FUNGIDB
CAZyme Information: CAH02555.1
You are here: Home > Sequence: CAH02555.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
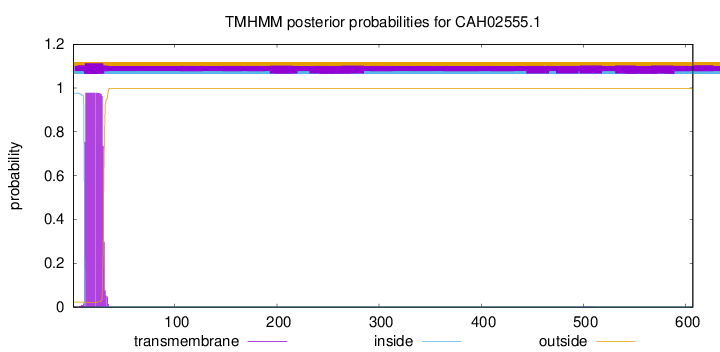
TMHMM annotations
Basic Information help
| Species | Kluyveromyces lactis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Saccharomycetes; ; Saccharomycetaceae; Kluyveromyces; Kluyveromyces lactis | |||||||||||
| CAZyme ID | CAH02555.1 | |||||||||||
| CAZy Family | GT58 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH125 | 128 | 575 | 2.5e-158 | 0.9950248756218906 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 399660 | Glyco_hydro_125 | 0.0 | 128 | 575 | 1 | 415 | Metal-independent alpha-mannosidase (GH125). This family, which contains bacterial and fungal glycoside hydrolases, is also known as GH125. They function as metal-independent alpha-mannosidases, with specificity for alpha-1,6-linked non-reducing terminal mannose residues. Structurally this family is part of the 6 hairpin glycosidase superfamily. |
| 226068 | COG3538 | 2.09e-152 | 107 | 583 | 2 | 427 | Meiotically up-regulated gene 157 (Mug157) protein (function unknown) [Function unknown]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 607 | 1 | 607 | |
| 0.0 | 1 | 607 | 1 | 607 | |
| 0.0 | 1 | 607 | 1 | 607 | |
| 0.0 | 4 | 605 | 4 | 603 | |
| 0.0 | 4 | 605 | 4 | 603 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.76e-97 | 102 | 578 | 43 | 468 | Crystal structure of BT3781 protein from Bacteroides thetaiotaomicron, Northeast Structural Genomics Target BtR58 [Bacteroides thetaiotaomicron VPI-5482],2P0V_B Crystal structure of BT3781 protein from Bacteroides thetaiotaomicron, Northeast Structural Genomics Target BtR58 [Bacteroides thetaiotaomicron VPI-5482] |
|
| 7.00e-95 | 102 | 578 | 23 | 448 | Crystal structure of an exo-alpha-1,6-mannosidase (bacova_03626) from bacteroides ovatus at 1.70 a resolution [Bacteroides ovatus ATCC 8483],3ON6_B Crystal structure of an exo-alpha-1,6-mannosidase (bacova_03626) from bacteroides ovatus at 1.70 a resolution [Bacteroides ovatus ATCC 8483] |
|
| 5.76e-94 | 102 | 578 | 21 | 450 | Crystal structure of an exo-alpha-1,6-mannosidase (bacova_03347) from bacteroides ovatus at 1.60 a resolution [Bacteroides ovatus ATCC 8483],3P2C_B Crystal structure of an exo-alpha-1,6-mannosidase (bacova_03347) from bacteroides ovatus at 1.60 a resolution [Bacteroides ovatus ATCC 8483] |
|
| 8.02e-74 | 113 | 576 | 10 | 420 | Crystal structure of GH125 1,6-alpha-mannosidase from Clostridium perfringens in complex with mannoimidazole [Clostridium perfringens str. 13],6RQK_B Crystal structure of GH125 1,6-alpha-mannosidase from Clostridium perfringens in complex with mannoimidazole [Clostridium perfringens str. 13] |
|
| 4.32e-73 | 113 | 576 | 10 | 420 | Crystal structure of GH125 1,6-alpha-mannosidase mutant from Clostridium perfringens in complex with 1,6-alpha-mannobiose [Clostridium perfringens str. 13],5M7Y_A Crystal structure of GH125 1,6-alpha-mannosidase mutant from Clostridium perfringens in complex with 1,6-alpha-mannotriose [Clostridium perfringens str. 13] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.72e-162 | 74 | 586 | 25 | 504 | Meiotically up-regulated gene 157 protein OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=mug157 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000023 | 0.000006 |