You are browsing environment: FUNGIDB
CAZyme Information: CAGL0D06490g-T-p1
You are here: Home > Sequence: CAGL0D06490g-T-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | [Candida] glabrata | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Saccharomycetes; ; Debaryomycetaceae; Candida; [Candida] glabrata | |||||||||||
| CAZyme ID | CAGL0D06490g-T-p1 | |||||||||||
| CAZy Family | GH16 | |||||||||||
| CAZyme Description | Ortholog(s) have chitin deacetylase activity and role in ascospore wall assembly | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.5.1.41:8 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE4 | 164 | 275 | 5.6e-24 | 0.8692307692307693 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 200576 | CE4_MrCDA_like | 2.64e-69 | 161 | 329 | 1 | 173 | Catalytic NodB homology domain of Mucor rouxii chitin deacetylase and similar proteins. This family is represented by the chitin deacetylase (MrCDA, EC 3.5.1.41) encoded from the fungus Mucor rouxii (also known as Amylomyces rouxii). MrCDA is an acidic glycoprotein with a very stringent specificity for beta1-4-linked N-acetylglucosamine homopolymers. It requires at least four residues (chitotetraose) for catalysis, and can achieve extensive deacetylation on chitin polymers. MrCDA shows high sequence similarity to Colletotrichum lindemuthianum chitin deacetylase (endo-chitin de-N-acetylase, ClCDA), which consists of a single catalytic domain similar to the deformed (beta/alpha)8 barrel fold adopted by the carbohydrate esterase 4 (CE4) superfamily, which encompasses a mononuclear metalloenzyme employing a conserved His-His-Asp zinc-binding triad closely associated with the conserved catalytic base (aspartic acid) and acid (histidine) to carry out acid/base catalysis. The family also includes some uncharacterized eukaryotic and bacterial homologs of MrCDA. |
| 213022 | CE4_NodB_like_6s_7s | 4.84e-48 | 166 | 333 | 6 | 167 | Catalytic NodB homology domain of rhizobial NodB-like proteins. This family belongs to the large and functionally diverse carbohydrate esterase 4 (CE4) superfamily, whose members show strong sequence similarity with some variability due to their distinct carbohydrate substrates. It includes many rhizobial NodB chitooligosaccharide N-deacetylase (EC 3.5.1.-)-like proteins, mainly from bacteria and eukaryotes, such as chitin deacetylases (EC 3.5.1.41), bacterial peptidoglycan N-acetylglucosamine deacetylases (EC 3.5.1.-), and acetylxylan esterases (EC 3.1.1.72), which catalyze the N- or O-deacetylation of substrates such as acetylated chitin, peptidoglycan, and acetylated xylan. All members of this family contain a catalytic NodB homology domain with the same overall topology and a deformed (beta/alpha)8 barrel fold with 6- or 7 strands. Their catalytic activity is dependent on the presence of a divalent cation, preferably cobalt or zinc, and they employ a conserved His-His-Asp zinc-binding triad closely associated with the conserved catalytic base (aspartic acid) and acid (histidine) to carry out acid/base catalysis. Several family members show diversity both in metal ion specificities and in the residues that coordinate the metal. |
| 200575 | CE4_ClCDA_like | 3.01e-40 | 166 | 322 | 13 | 175 | Catalytic NodB homology domain of Colletotrichum lindemuthianum chitin deacetylase and similar proteins. This family is represented by the chitin deacetylase (endo-chitin de-N-acetylase, ClCDA, EC 3.5.1.41) from Colletotrichum lindemuthianum (also known as Glomerella lindemuthiana), which is a member of the carbohydrate esterase 4 (CE4) superfamily. ClCDA catalyzes the hydrolysis of N-acetamido groups of N-acetyl-D-glucosamine residues in chitin, converting it to chitosan in fungal cell walls. It consists of a single catalytic domain similar to the deformed (alpha/beta)8 barrel fold adopted by other CE4 esterases, which encompasses a mononuclear metalloenzyme employing a conserved His-His-Asp zinc-binding triad closely associated with the conserved catalytic base (aspartic acid) and acid (histidine), to carry out acid/base catalysis. It possesses a highly conserved substrate-binding groove, with subtle alterations that influence substrate specificity and subsite affinity. Unlike its bacterial homologs, ClCDA contains two intramolecular disulfide bonds that may add stability to this secreted protein. The family also includes many uncharacterized deacetylases and hypothetical proteins mainly from eukaryotes, which show high sequence similarity to ClCDA. |
| 396211 | Polysacc_deac_1 | 1.07e-39 | 158 | 275 | 4 | 124 | Polysaccharide deacetylase. This domain is found in polysaccharide deacetylase. This family of polysaccharide deacetylases includes NodB (nodulation protein B from Rhizobium) which is a chitooligosaccharide deacetylase. It also includes chitin deacetylase from yeast, and endoxylanases which hydrolyzes glucosidic bonds in xylan. |
| 223798 | CDA1 | 2.41e-37 | 95 | 333 | 3 | 240 | Peptidoglycan/xylan/chitin deacetylase, PgdA/CDA1 family [Carbohydrate transport and metabolism, Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.59e-268 | 1 | 354 | 1 | 354 | |
| 1.59e-268 | 1 | 354 | 1 | 354 | |
| 2.16e-268 | 1 | 354 | 9 | 362 | |
| 1.25e-267 | 1 | 354 | 9 | 362 | |
| 1.25e-267 | 1 | 354 | 9 | 362 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4.48e-23 | 166 | 345 | 45 | 228 | Chain A, Aspergillus niger contig An12c0130, genomic contig [Aspergillus niger CBS 513.88] |
|
| 1.71e-21 | 166 | 329 | 15 | 172 | Structure of peptidoglycan deacetylase PdaC from Bacillus subtilis [Bacillus subtilis subsp. subtilis str. 168],6H8L_B Structure of peptidoglycan deacetylase PdaC from Bacillus subtilis [Bacillus subtilis subsp. subtilis str. 168] |
|
| 4.87e-21 | 166 | 322 | 122 | 275 | Chain A, Predicted xylanase/chitin deacetylase [Caldanaerobacter subterraneus subsp. tengcongensis MB4] |
|
| 3.92e-20 | 166 | 349 | 30 | 209 | T48 deacetylase [Arthrobacter sp. AW19M34-1],5LGC_A T48 deacetylase with substrate [Arthrobacter sp. AW19M34-1] |
|
| 2.84e-19 | 166 | 322 | 10 | 161 | Family 4 carbohydrate esterase from Streptomyces lividans in complex with acetate [Streptomyces lividans],2CC0_B Family 4 carbohydrate esterase from Streptomyces lividans in complex with acetate [Streptomyces lividans] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.60e-128 | 85 | 353 | 42 | 310 | Chitin deacetylase 2 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=CDA2 PE=1 SV=1 |
|
| 7.42e-117 | 91 | 353 | 38 | 300 | Chitin deacetylase 1 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=CDA1 PE=1 SV=1 |
|
| 2.02e-31 | 120 | 327 | 112 | 342 | Chitin deacetylase 1 OS=Cryptococcus neoformans var. grubii serotype A (strain H99 / ATCC 208821 / CBS 10515 / FGSC 9487) OX=235443 GN=CDA1 PE=1 SV=1 |
|
| 4.41e-30 | 142 | 286 | 103 | 256 | Chitin deacetylase 3 OS=Cryptococcus neoformans var. neoformans serotype D (strain JEC21 / ATCC MYA-565) OX=214684 GN=CDA3 PE=2 SV=1 |
|
| 6.09e-30 | 142 | 286 | 103 | 256 | Chitin deacetylase 3 OS=Cryptococcus neoformans var. neoformans serotype D (strain B-3501A) OX=283643 GN=CDA3 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
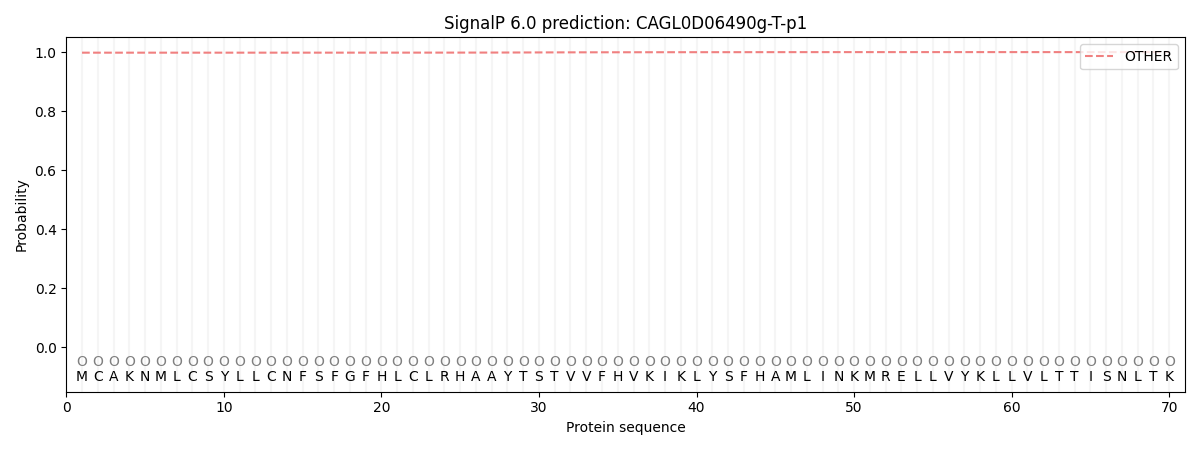
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.998281 | 0.001730 |
