You are browsing environment: FUNGIDB
CAZyme Information: An11g05850-T-p1
You are here: Home > Sequence: An11g05850-T-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus niger | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus niger | |||||||||||
| CAZyme ID | An11g05850-T-p1 | |||||||||||
| CAZy Family | GH43 | |||||||||||
| CAZyme Description | Has domain(s) with predicted UDP-N-acetylmuramate dehydrogenase activity, flavin adenine dinucleotide binding, oxidoreductase activity, acting on CH-OH group of donors activity and role in oxidation-reduction process | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA7 | 37 | 198 | 2e-29 | 0.35807860262008734 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 223354 | GlcD | 2.11e-07 | 39 | 178 | 25 | 167 | FAD/FMN-containing dehydrogenase [Energy production and conversion]. |
| 223882 | MurB | 1.77e-04 | 47 | 191 | 22 | 162 | UDP-N-acetylenolpyruvoylglucosamine reductase [Cell wall/membrane/envelope biogenesis]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 4.70e-10 | 1 | 198 | 17 | 218 | |
| 8.25e-10 | 1 | 198 | 17 | 218 | |
| 2.55e-09 | 36 | 198 | 52 | 218 | |
| 6.47e-09 | 1 | 197 | 46 | 265 | |
| 7.44e-09 | 26 | 198 | 27 | 205 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.17e-15 | 22 | 492 | 15 | 453 | Crystal structure of 6-hydoxy-D-nicotine oxidase from Arthrobacter nicotinovorans. Crystal Form 3 (P1) [Paenarthrobacter nicotinovorans],2BVF_B Crystal structure of 6-hydoxy-D-nicotine oxidase from Arthrobacter nicotinovorans. Crystal Form 3 (P1) [Paenarthrobacter nicotinovorans],2BVG_A Crystal structure of 6-hydoxy-D-nicotine oxidase from Arthrobacter nicotinovorans. Crystal Form 1 (P21) [Paenarthrobacter nicotinovorans],2BVG_B Crystal structure of 6-hydoxy-D-nicotine oxidase from Arthrobacter nicotinovorans. Crystal Form 1 (P21) [Paenarthrobacter nicotinovorans],2BVG_C Crystal structure of 6-hydoxy-D-nicotine oxidase from Arthrobacter nicotinovorans. Crystal Form 1 (P21) [Paenarthrobacter nicotinovorans],2BVG_D Crystal structure of 6-hydoxy-D-nicotine oxidase from Arthrobacter nicotinovorans. Crystal Form 1 (P21) [Paenarthrobacter nicotinovorans],2BVH_A Crystal structure of 6-hydoxy-D-nicotine oxidase from Arthrobacter nicotinovorans. Crystal Form 2 (P21) [Paenarthrobacter nicotinovorans],2BVH_B Crystal structure of 6-hydoxy-D-nicotine oxidase from Arthrobacter nicotinovorans. Crystal Form 2 (P21) [Paenarthrobacter nicotinovorans],2BVH_C Crystal structure of 6-hydoxy-D-nicotine oxidase from Arthrobacter nicotinovorans. Crystal Form 2 (P21) [Paenarthrobacter nicotinovorans],2BVH_D Crystal structure of 6-hydoxy-D-nicotine oxidase from Arthrobacter nicotinovorans. Crystal Form 2 (P21) [Paenarthrobacter nicotinovorans] |
|
| 3.39e-13 | 32 | 199 | 55 | 229 | Berberine bridge enzyme G164A variant, a reticuline dehydrogenase [Eschscholzia californica],4PZF_B Berberine bridge enzyme G164A variant, a reticuline dehydrogenase [Eschscholzia californica],4PZF_C Berberine bridge enzyme G164A variant, a reticuline dehydrogenase [Eschscholzia californica],4PZF_D Berberine bridge enzyme G164A variant, a reticuline dehydrogenase [Eschscholzia californica] |
|
| 4.31e-13 | 5 | 195 | 26 | 208 | Crystal structure of tetrahydrocannabinolic acid synthase from Cannabis sativa [Cannabis sativa] |
|
| 5.73e-13 | 32 | 199 | 36 | 210 | Structure of berberine bridge enzyme, H174A variant in complex with (S)-reticuline [Eschscholzia californica] |
|
| 7.20e-13 | 32 | 199 | 30 | 204 | Chain A, Reticuline oxidase [Eschscholzia californica] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.04e-15 | 38 | 269 | 70 | 286 | FAD-dependent monooxygenase tpcD OS=Cochliobolus heterostrophus (strain C5 / ATCC 48332 / race O) OX=701091 GN=tpcD PE=1 SV=1 |
|
| 1.48e-14 | 22 | 492 | 14 | 452 | (R)-6-hydroxynicotine oxidase OS=Paenarthrobacter nicotinovorans OX=29320 GN=6-hdno PE=1 SV=2 |
|
| 9.41e-14 | 2 | 268 | 18 | 269 | Uncharacterized FAD-linked oxidoreductase ARB_02372 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_02372 PE=1 SV=1 |
|
| 1.65e-12 | 21 | 198 | 41 | 223 | FAD-dependent monooxygenase prx3 OS=Penicillium roqueforti (strain FM164) OX=1365484 GN=prx3 PE=3 SV=1 |
|
| 1.65e-12 | 21 | 198 | 41 | 223 | FAD-dependent monooxygenase prx3 OS=Penicillium roqueforti OX=5082 GN=prx3 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
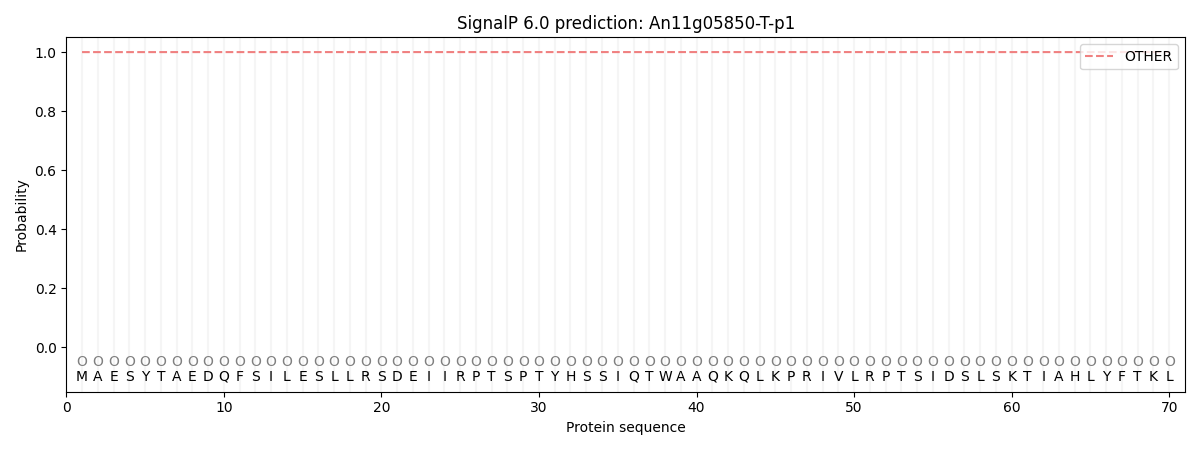
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000048 | 0.000000 |
