You are browsing environment: FUNGIDB
CAZyme Information: An04g10320-T-p1
You are here: Home > Sequence: An04g10320-T-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
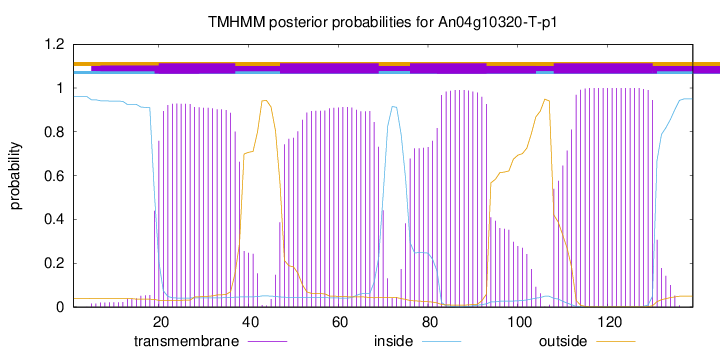
TMHMM annotations
Basic Information help
| Species | Aspergillus niger | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus niger | |||||||||||
| CAZyme ID | An04g10320-T-p1 | |||||||||||
| CAZy Family | GH13 | |||||||||||
| CAZyme Description | protein of unknown function | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA6 | 20 | 119 | 8.5e-34 | 0.5076923076923077 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 130816 | flav_wrbA | 6.35e-25 | 20 | 122 | 69 | 169 | NAD(P)H:quinone oxidoreductase, type IV. This model represents a protein, WrbA, related to and slightly larger than flavodoxin. It was just shown, in E. coli and Archaeoglobus fulgidus (and previously for some eukaryotic homologs) to act as fourth type of NAD(P)H:quinone oxidoreductase. In E. coli, this protein was earlier reported to be produced during stationary phase, bind to the trp repressor, and make trp operon repression more efficient. WrbA does not interact with the trp operator by itself. Members are found in species in which homologs of the E. coli trp operon repressor TrpR (SP:P03032) are not detected. [Energy metabolism, Electron transport] |
| 179647 | PRK03767 | 1.27e-24 | 20 | 117 | 70 | 165 | NAD(P)H:quinone oxidoreductase; Provisional |
| 223728 | WrbA | 6.97e-16 | 20 | 125 | 76 | 181 | Multimeric flavodoxin WrbA [Energy production and conversion]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 5.09e-97 | 1 | 139 | 1 | 139 | |
| 1.44e-47 | 20 | 120 | 70 | 170 | |
| 1.44e-47 | 20 | 120 | 70 | 170 | |
| 2.05e-47 | 20 | 120 | 70 | 170 | |
| 2.05e-47 | 20 | 120 | 70 | 170 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.95e-34 | 20 | 115 | 69 | 163 | The structure of Pst2p from Saccharomyces cerevisiae [Saccharomyces cerevisiae S288C],5MP4_B The structure of Pst2p from Saccharomyces cerevisiae [Saccharomyces cerevisiae S288C],5MP4_C The structure of Pst2p from Saccharomyces cerevisiae [Saccharomyces cerevisiae S288C],5MP4_D The structure of Pst2p from Saccharomyces cerevisiae [Saccharomyces cerevisiae S288C],5MP4_E The structure of Pst2p from Saccharomyces cerevisiae [Saccharomyces cerevisiae S288C],5MP4_F The structure of Pst2p from Saccharomyces cerevisiae [Saccharomyces cerevisiae S288C],5MP4_G The structure of Pst2p from Saccharomyces cerevisiae [Saccharomyces cerevisiae S288C],5MP4_H The structure of Pst2p from Saccharomyces cerevisiae [Saccharomyces cerevisiae S288C] |
|
| 5.08e-18 | 20 | 115 | 68 | 160 | X-ray structure of E.coli Wrba in complex with FMN at 1.2 A resolution [Escherichia coli],3ZHO_B X-ray structure of E.coli Wrba in complex with FMN at 1.2 A resolution [Escherichia coli],4YQE_A Crystal structure of E. coli WrbA in complex with benzoquinone [Escherichia coli K-12],4YQE_B Crystal structure of E. coli WrbA in complex with benzoquinone [Escherichia coli K-12],5F12_A WrbA in complex with FMN under crystallization conditions of WrbA-FMN-BQ structure (4YQE) [Escherichia coli K-12],5F12_B WrbA in complex with FMN under crystallization conditions of WrbA-FMN-BQ structure (4YQE) [Escherichia coli K-12] |
|
| 5.08e-18 | 20 | 115 | 68 | 160 | High resolution structure of E.coli WrbA with FMN [Escherichia coli APEC O1],4DY4_C High resolution structure of E.coli WrbA with FMN [Escherichia coli APEC O1] |
|
| 5.18e-18 | 20 | 115 | 69 | 161 | Crystal structure of E. coli WrbA in complex with FMN [Escherichia coli],2R96_C Crystal structure of E. coli WrbA in complex with FMN [Escherichia coli],2R97_A Crystal structure of E. coli WrbA in complex with FMN [Escherichia coli],2R97_C Crystal structure of E. coli WrbA in complex with FMN [Escherichia coli],2RG1_A Crystal structure of E. coli WrbA apoprotein [Escherichia coli],2RG1_B Crystal structure of E. coli WrbA apoprotein [Escherichia coli],3B6I_A WrbA from Escherichia coli, native structure [Escherichia coli K-12],3B6I_B WrbA from Escherichia coli, native structure [Escherichia coli K-12],3B6J_A WrbA from Escherichia coli, NADH complex [Escherichia coli K-12],3B6J_B WrbA from Escherichia coli, NADH complex [Escherichia coli K-12],3B6K_A WrbA from Escherichia coli, Benzoquinone complex [Escherichia coli K-12],3B6K_B WrbA from Escherichia coli, Benzoquinone complex [Escherichia coli K-12],3B6M_A WrbA from Escherichia coli, second crystal form [Escherichia coli K-12],3B6M_B WrbA from Escherichia coli, second crystal form [Escherichia coli K-12] |
|
| 2.52e-16 | 21 | 117 | 76 | 170 | Crystal structure of native PnpB [Pseudomonas sp. WBC-3],4LA4_B Crystal structure of native PnpB [Pseudomonas sp. WBC-3],4LAF_A Crystal structure of PnpB complex with FMN [Pseudomonas sp. WBC-3],4LAF_B Crystal structure of PnpB complex with FMN [Pseudomonas sp. WBC-3],4LAF_C Crystal structure of PnpB complex with FMN [Pseudomonas sp. WBC-3],4LAF_D Crystal structure of PnpB complex with FMN [Pseudomonas sp. WBC-3] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.86e-37 | 20 | 122 | 72 | 174 | Minor allergen Alt a 7 OS=Alternaria alternata OX=5599 GN=ALTA7 PE=1 SV=1 |
|
| 1.04e-34 | 29 | 122 | 81 | 174 | Minor allergen Cla h 7 OS=Davidiella tassiana OX=29918 GN=CLAH7 PE=1 SV=1 |
|
| 1.00e-33 | 20 | 115 | 69 | 163 | Protoplast secreted protein 2 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=PST2 PE=1 SV=1 |
|
| 2.38e-30 | 20 | 115 | 70 | 165 | Flavoprotein-like protein YCP4 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=YCP4 PE=1 SV=1 |
|
| 2.26e-27 | 20 | 115 | 72 | 167 | P25 protein OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=obr1 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.999957 | 0.000050 |