You are browsing environment: FUNGIDB
CAZyme Information: An01g04560-T-p1
You are here: Home > Sequence: An01g04560-T-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus niger | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus niger | |||||||||||
| CAZyme ID | An01g04560-T-p1 | |||||||||||
| CAZy Family | AA1 | |||||||||||
| CAZyme Description | Beta-glucanase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.73:4 | 3.2.1.6:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH16 | 68 | 297 | 2.1e-94 | 0.9737991266375546 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 185690 | GH16_fungal_Lam16A_glucanase | 2.33e-150 | 25 | 318 | 1 | 293 | fungal 1,3(4)-beta-D-glucanases, similar to Phanerochaete chrysosporium laminarinase 16A. Group of fungal 1,3(4)-beta-D-glucanases, similar to Phanerochaete chrysosporium laminarinase 16A. Lam16A belongs to the 'nonspecific' 1,3(4)-beta-glucanase subfamily, although beta-1,6 branching and beta-1,4 bonds specifically define where Lam16A hydrolyzes its substrates, like curdlan (beta-1,3-glucan), lichenin (beta-1,3-1,4-mixed linkage glucan), and laminarin (beta-1,6-branched-1,3-glucan). |
| 185693 | GH16_laminarinase_like | 1.69e-11 | 113 | 283 | 88 | 220 | Laminarinase, member of the glycosyl hydrolase family 16. Laminarinase, also known as glucan endo-1,3-beta-D-glucosidase, is a glycosyl hydrolase family 16 member that hydrolyzes 1,3-beta-D-glucosidic linkages in 1,3-beta-D-glucans such as laminarins, curdlans, paramylons, and pachymans, with very limited action on mixed-link (1,3-1,4-)-beta-D-glucans. |
| 185691 | GH16_Strep_laminarinase_like | 8.15e-06 | 119 | 154 | 106 | 148 | Streptomyces laminarinase-like, member of glycosyl hydrolase family 16. Proteins similar to Streptomyces sioyaensis beta-1,3-glucanase (laminarinase) present in Actinomycetales as well as Peziomycotina. Laminarinases belong to glycosyl hydrolase family 16 and hydrolyze the glycosidic bond of the 1,3-beta-linked glucan, a major component of fungal and plant cell walls and the structural and storage polysaccharides (laminarin) of marine macro-algae. Members of the GH16 family have a conserved jelly roll fold with an active site channel. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 3.14e-278 | 1 | 432 | 1 | 432 | |
| 1.05e-246 | 5 | 432 | 1 | 432 | |
| 1.05e-246 | 5 | 432 | 1 | 432 | |
| 9.65e-153 | 5 | 325 | 1 | 322 | |
| 4.66e-150 | 7 | 319 | 2 | 309 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.62e-117 | 26 | 317 | 2 | 295 | The complex structure of PtLic16A with cellobiose [Paecilomyces sp. 'thermophila'],3WDU_B The complex structure of PtLic16A with cellobiose [Paecilomyces sp. 'thermophila'],3WDU_C The complex structure of PtLic16A with cellobiose [Paecilomyces sp. 'thermophila'],3WDU_D The complex structure of PtLic16A with cellobiose [Paecilomyces sp. 'thermophila'] |
|
| 9.95e-117 | 26 | 317 | 3 | 296 | The apo-form structure of PtLic16A from Paecilomyces thermophila [Paecilomyces sp. 'thermophila'],3WDT_B The apo-form structure of PtLic16A from Paecilomyces thermophila [Paecilomyces sp. 'thermophila'],3WDT_C The apo-form structure of PtLic16A from Paecilomyces thermophila [Paecilomyces sp. 'thermophila'],3WDT_D The apo-form structure of PtLic16A from Paecilomyces thermophila [Paecilomyces sp. 'thermophila'],3WDV_A The complex structure of PtLic16A with cellotetraose [Paecilomyces sp. 'thermophila'],3WDV_B The complex structure of PtLic16A with cellotetraose [Paecilomyces sp. 'thermophila'],3WDV_C The complex structure of PtLic16A with cellotetraose [Paecilomyces sp. 'thermophila'],3WDV_D The complex structure of PtLic16A with cellotetraose [Paecilomyces sp. 'thermophila'] |
|
| 7.51e-116 | 26 | 317 | 1 | 294 | The complex structure of E113A with cellotetraose [Paecilomyces sp. 'thermophila'],3WDY_B The complex structure of E113A with cellotetraose [Paecilomyces sp. 'thermophila'],3WDY_C The complex structure of E113A with cellotetraose [Paecilomyces sp. 'thermophila'],3WDY_D The complex structure of E113A with cellotetraose [Paecilomyces sp. 'thermophila'] |
|
| 8.04e-116 | 26 | 317 | 3 | 296 | The apo-form structure of E113A from Paecilomyces thermophila [Paecilomyces sp. 'thermophila'],3WDW_B The apo-form structure of E113A from Paecilomyces thermophila [Paecilomyces sp. 'thermophila'],3WDX_A The complex structure of E113A with glucotriose [Paecilomyces sp. 'thermophila'],3WDX_B The complex structure of E113A with glucotriose [Paecilomyces sp. 'thermophila'] |
|
| 3.40e-96 | 25 | 317 | 2 | 297 | Crystal structure and characterization an elongating GH family 16 beta-1,3-glucosyltransferase [Paecilomyces sp. 'thermophila'],5JVV_B Crystal structure and characterization an elongating GH family 16 beta-1,3-glucosyltransferase [Paecilomyces sp. 'thermophila'] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.63e-109 | 5 | 318 | 1 | 315 | Endo-1,3(4)-beta-glucanase ARB_04519 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_04519 PE=3 SV=1 |
|
| 6.32e-93 | 26 | 318 | 129 | 418 | Probable glycosidase C21B10.07 OS=Schizosaccharomyces pombe (strain 972 / ATCC 24843) OX=284812 GN=SPBC21B10.07 PE=3 SV=1 |
|
| 9.98e-93 | 8 | 318 | 15 | 327 | Probable endo-1,3(4)-beta-glucanase AFUB_029980 OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=AFUB_029980 PE=3 SV=1 |
|
| 9.98e-93 | 8 | 318 | 15 | 327 | Probable endo-1,3(4)-beta-glucanase AFUA_2G14360 OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=AFUA_2G14360 PE=3 SV=1 |
|
| 5.35e-92 | 10 | 318 | 17 | 327 | Probable endo-1,3(4)-beta-glucanase NFIA_089530 OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=NFIA_089530 PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
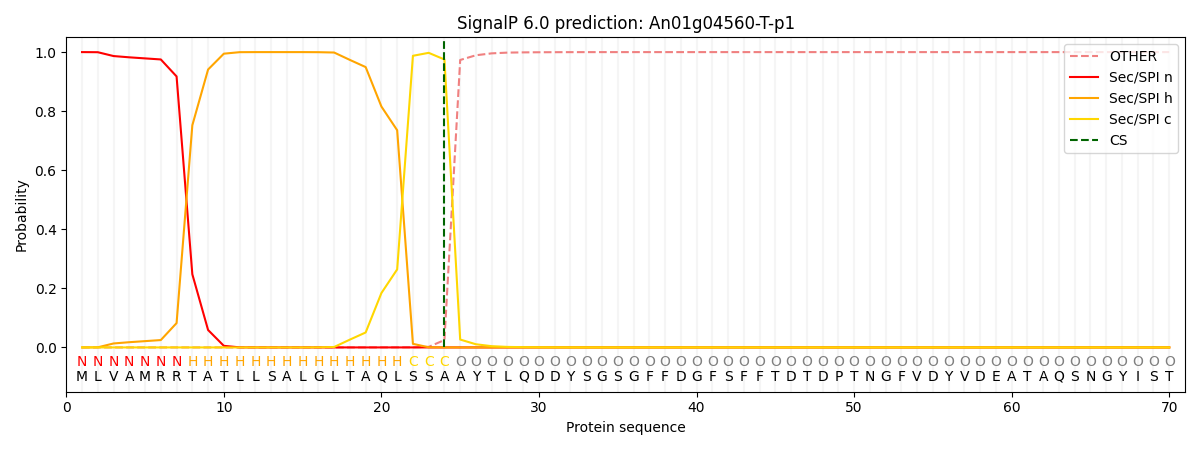
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000231 | 0.999756 | CS pos: 24-25. Pr: 0.9750 |
