You are browsing environment: FUNGIDB
CAZyme Information: Afu3g00470-T-p1
You are here: Home > Sequence: Afu3g00470-T-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus fumigatus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus fumigatus | |||||||||||
| CAZyme ID | Afu3g00470-T-p1 | |||||||||||
| CAZy Family | GH15 | |||||||||||
| CAZyme Description | Has domain(s) with predicted cellulose binding, hydrolase activity, hydrolyzing O-glycosyl compounds activity, role in carbohydrate metabolic process and extracellular region localization | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.8:67 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH11 | 43 | 221 | 1.9e-74 | 0.9943502824858758 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 395367 | Glyco_hydro_11 | 7.24e-106 | 43 | 219 | 1 | 175 | Glycosyl hydrolases family 11. |
| 197593 | fCBD | 1.99e-10 | 254 | 286 | 1 | 34 | Fungal-type cellulose-binding domain. Small four-cysteine cellulose-binding domain of fungi |
| 395595 | CBM_1 | 1.17e-09 | 255 | 282 | 1 | 29 | Fungal cellulose binding domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.05e-163 | 1 | 229 | 1 | 229 | |
| 1.04e-139 | 1 | 286 | 1 | 310 | |
| 1.04e-139 | 1 | 286 | 1 | 310 | |
| 1.04e-139 | 1 | 286 | 1 | 310 | |
| 1.32e-131 | 1 | 230 | 1 | 230 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.32e-103 | 32 | 223 | 1 | 190 | Crystal structure of family 11 xylanase in complex with inhibitor (XIP-I) [Talaromyces funiculosus] |
|
| 1.67e-103 | 28 | 223 | 14 | 207 | Xylanase 11C from Talaromyces cellulolyticus (formerly known as Acremonium cellulolyticus) [Talaromyces funiculosus],3WP3_B Xylanase 11C from Talaromyces cellulolyticus (formerly known as Acremonium cellulolyticus) [Talaromyces funiculosus] |
|
| 9.60e-99 | 34 | 223 | 3 | 190 | Chain A, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_B Chain B, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_C Chain C, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_D Chain D, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_E Chain E, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_F Chain F, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_G Chain G, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_H Chain H, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_I Chain I, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_J Chain J, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_K Chain K, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612],5HXV_L Chain L, Endo-1,4-beta-xylanase [Talaromyces cellulolyticus CF-2612] |
|
| 4.56e-87 | 39 | 223 | 6 | 189 | Chain A, Endo-1,4-beta-xylanase 2 [Trichoderma reesei],4XQ4_A Chain A, Endo-1,4-beta-xylanase 2 [Trichoderma reesei],4XQ4_B Chain B, Endo-1,4-beta-xylanase 2 [Trichoderma reesei],6JXL_A Crystal Structures of Endo-beta-1,4-xylanase II Complexed with Xylotriose [Trichoderma reesei RUT C-30],6K9O_A Crystal Structure Analysis of Protein [Trichoderma reesei RUT C-30],6KWD_A Crystal Structure Analysis of Endo-beta-1,4-Xylanase II Complexed with Xylotriose [Trichoderma reesei RUT C-30] |
|
| 4.72e-87 | 39 | 223 | 7 | 190 | Chain A, Endo-1,4-beta-xylanase 2 [Trichoderma reesei RUT C-30],6KW9_A Crystal Structure Analysis of Endo-beta-1,4-xylanase II Complexed with Xylotriose [Trichoderma reesei RUT C-30],6KWF_A Crystal Structure Analysis of Endo-beta-1,4-xylanase II Complexed with Xylotriose [Trichoderma reesei RUT C-30] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.85e-140 | 1 | 286 | 1 | 310 | Endo-1,4-beta-xylanase B OS=Penicillium oxalicum OX=69781 GN=xynB PE=1 SV=1 |
|
| 3.89e-116 | 1 | 286 | 1 | 290 | Endo-1,4-beta-xylanase B OS=Phanerodontia chrysosporium OX=2822231 GN=xynB PE=1 SV=1 |
|
| 1.03e-110 | 1 | 223 | 1 | 216 | Endo-1,4-beta-xylanase 2 OS=Rhizopus oryzae OX=64495 GN=xyn2 PE=1 SV=1 |
|
| 4.51e-104 | 28 | 223 | 30 | 223 | Endo-1,4-beta-xylanase C OS=Talaromyces funiculosus OX=28572 GN=xynC PE=1 SV=1 |
|
| 3.78e-99 | 1 | 223 | 1 | 217 | Endo-1,4-beta-xylanase A OS=Penicillium citrinum OX=5077 GN=xynA PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
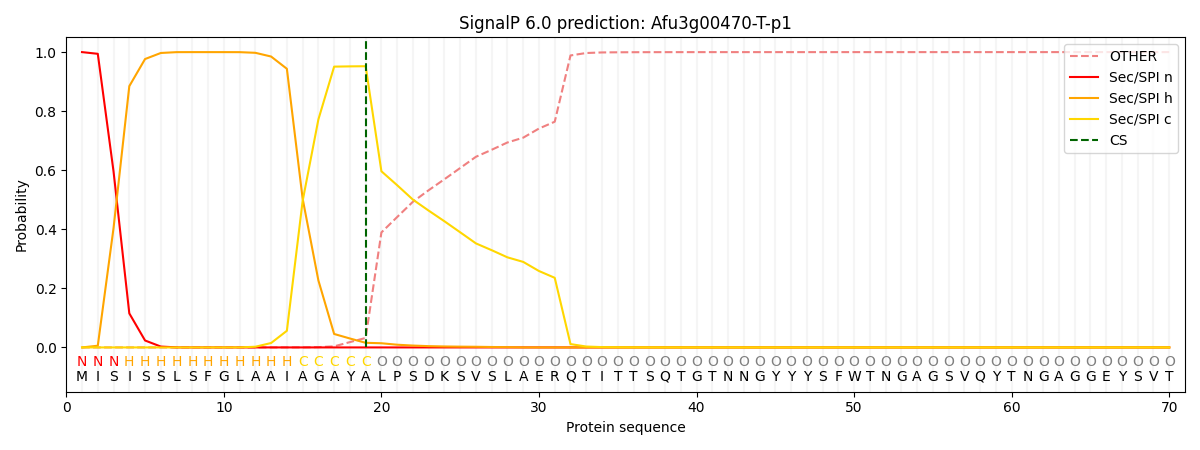
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000350 | 0.999604 | CS pos: 19-20. Pr: 0.9522 |
