You are browsing environment: FUNGIDB
CAZyme Information: AWU77555.1
You are here: Home > Sequence: AWU77555.1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
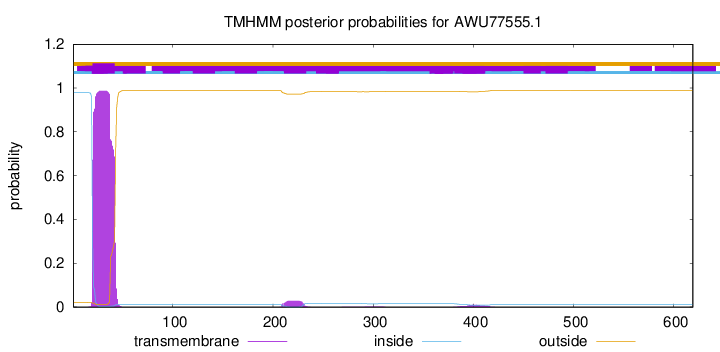
TMHMM annotations
Basic Information help
| Species | Pichia kudriavzevii | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Saccharomycetes; ; Pichiaceae; Pichia; Pichia kudriavzevii | |||||||||||
| CAZyme ID | AWU77555.1 | |||||||||||
| CAZy Family | GT58 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT71 | 213 | 470 | 1.9e-65 | 0.9924242424242424 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 402574 | Mannosyl_trans3 | 8.57e-56 | 214 | 448 | 3 | 248 | Mannosyltransferase putative. This family is conserved in fungi. Several members are annotated as being alpha-1,3-mannosyltransferase but this could not be confirmed. |
| 133018 | GT8_Glycogenin | 9.42e-05 | 223 | 319 | 12 | 110 | Glycogenin belongs the GT 8 family and initiates the biosynthesis of glycogen. Glycogenin initiates the biosynthesis of glycogen by incorporating glucose residues through a self-glucosylation reaction at a Tyr residue, and then acts as substrate for chain elongation by glycogen synthase and branching enzyme. It contains a conserved DxD motif and an N-terminal beta-alpha-beta Rossmann-like fold that are common to the nucleotide-binding domains of most glycosyltransferases. The DxD motif is essential for coordination of the catalytic divalent cation, most commonly Mn2+. Glycogenin can be classified as a retaining glycosyltransferase, based on the relative anomeric stereochemistry of the substrate and product in the reaction catalyzed. It is placed in glycosyltransferase family 8 which includes lipopolysaccharide glucose and galactose transferases and galactinol synthases. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 619 | 1 | 619 | |
| 2.18e-150 | 69 | 616 | 75 | 634 | |
| 3.79e-141 | 126 | 616 | 70 | 567 | |
| 2.59e-102 | 127 | 605 | 210 | 690 | |
| 2.22e-96 | 115 | 619 | 87 | 604 |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.43e-60 | 162 | 579 | 201 | 624 | Alpha-1,2-mannosyltransferase MNN21 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=MNN21 PE=3 SV=1 |
|
| 5.10e-59 | 40 | 616 | 84 | 706 | Alpha-1,2-mannosyltransferase MNN24 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=MNN24 PE=3 SV=1 |
|
| 1.47e-58 | 179 | 616 | 137 | 586 | Alpha-1,2-mannosyltransferase MNN2 OS=Saccharomyces cerevisiae (strain ATCC 204508 / S288c) OX=559292 GN=MNN2 PE=1 SV=1 |
|
| 3.19e-58 | 167 | 579 | 100 | 538 | Alpha-1,2-mannosyltransferase MNN5 OS=Saccharomyces cerevisiae (strain YJM789) OX=307796 GN=MNN5 PE=3 SV=1 |
|
| 1.22e-57 | 185 | 619 | 144 | 604 | Alpha-1,2-mannosyltransferase MNN23 OS=Candida albicans (strain SC5314 / ATCC MYA-2876) OX=237561 GN=MNN23 PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER

| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000031 | 0.000005 |