You are browsing environment: FUNGIDB
CAZyme Information: ATEG_09052-t26_1-p1
You are here: Home > Sequence: ATEG_09052-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus terreus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus terreus | |||||||||||
| CAZyme ID | ATEG_09052-t26_1-p1 | |||||||||||
| CAZy Family | GT21 | |||||||||||
| CAZyme Description | conserved hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.37:8 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH3 | 76 | 326 | 3.6e-68 | 0.9768518518518519 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 178629 | PLN03080 | 0.0 | 4 | 735 | 2 | 778 | Probable beta-xylosidase; Provisional |
| 185053 | PRK15098 | 4.06e-83 | 112 | 732 | 118 | 750 | beta-glucosidase BglX. |
| 224389 | BglX | 9.46e-56 | 56 | 423 | 35 | 365 | Periplasmic beta-glucosidase and related glycosidases [Carbohydrate transport and metabolism]. |
| 396478 | Glyco_hydro_3_C | 6.61e-47 | 398 | 621 | 1 | 216 | Glycosyl hydrolase family 3 C-terminal domain. This domain is involved in catalysis and may be involved in binding beta-glucan. This domain is found associated with pfam00933. |
| 395747 | Glyco_hydro_3 | 2.02e-33 | 78 | 355 | 63 | 313 | Glycosyl hydrolase family 3 N terminal domain. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 13 | 765 | 11 | 770 | |
| 0.0 | 13 | 765 | 11 | 770 | |
| 0.0 | 13 | 765 | 11 | 770 | |
| 0.0 | 26 | 765 | 45 | 791 | |
| 0.0 | 26 | 765 | 45 | 791 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 8.89e-305 | 29 | 758 | 2 | 742 | Chain A, xylan 1,4-beta-xylosidase [Phanerodontia chrysosporium] |
|
| 8.89e-305 | 29 | 758 | 2 | 742 | Chain A, xylan 1,4-beta-xylosidase [Phanerodontia chrysosporium] |
|
| 1.04e-261 | 29 | 756 | 29 | 764 | Chain A, BETA-XYLOSIDASE [Trichoderma reesei],5A7M_B Chain B, BETA-XYLOSIDASE [Trichoderma reesei] |
|
| 1.08e-261 | 29 | 756 | 29 | 764 | Chain A, BETA-XYLOSIDASE [Trichoderma reesei],5AE6_B Chain B, BETA-XYLOSIDASE [Trichoderma reesei] |
|
| 8.50e-258 | 29 | 757 | 29 | 763 | GH3 exo-beta-xylosidase (XlnD) [Aspergillus nidulans FGSC A4],6Q7I_B GH3 exo-beta-xylosidase (XlnD) [Aspergillus nidulans FGSC A4],6Q7J_A Chain A, Exo-1,4-beta-xylosidase xlnD [Aspergillus nidulans FGSC A4],6Q7J_B Chain B, Exo-1,4-beta-xylosidase xlnD [Aspergillus nidulans FGSC A4] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 1 | 765 | 1 | 765 | Probable exo-1,4-beta-xylosidase bxlB OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=bxlB PE=3 SV=1 |
|
| 0.0 | 13 | 760 | 13 | 769 | Probable exo-1,4-beta-xylosidase bxlB OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=bxlB PE=3 SV=1 |
|
| 0.0 | 26 | 765 | 45 | 791 | Probable exo-1,4-beta-xylosidase bxlB OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=bxlB PE=3 SV=1 |
|
| 0.0 | 13 | 760 | 13 | 769 | Probable exo-1,4-beta-xylosidase bxlB OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=bxlB PE=3 SV=1 |
|
| 0.0 | 13 | 765 | 11 | 770 | Probable exo-1,4-beta-xylosidase bxlB OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=bxlB PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
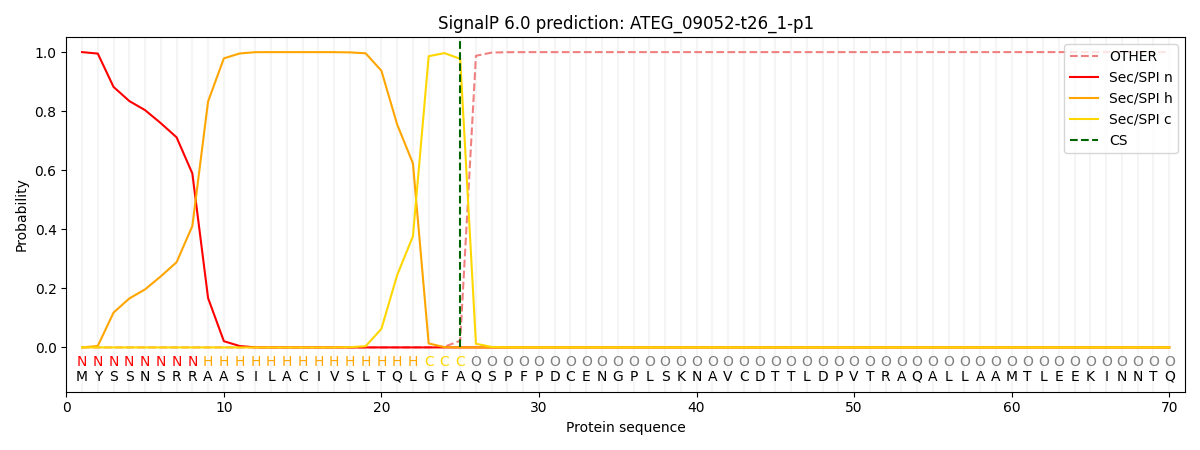
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000386 | 0.999581 | CS pos: 25-26. Pr: 0.9773 |
