You are browsing environment: FUNGIDB
CAZyme Information: ATEG_08684-t26_1-p1
You are here: Home > Sequence: ATEG_08684-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus terreus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus terreus | |||||||||||
| CAZyme ID | ATEG_08684-t26_1-p1 | |||||||||||
| CAZy Family | GT2 | |||||||||||
| CAZyme Description | conserved hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.25:3 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH2 | 6 | 681 | 1.9e-98 | 0.6316489361702128 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 225789 | LacZ | 3.12e-86 | 6 | 656 | 12 | 634 | Beta-galactosidase/beta-glucuronidase [Carbohydrate transport and metabolism]. |
| 396543 | Translin | 2.65e-53 | 901 | 1101 | 26 | 196 | Translin family. Members of this family include Translin, which interacts with DNA and forms a ring around the DNA. This family also includes human TSNAX, which was found to interact with translin with yeast two-hybrid screen. |
| 271350 | TRAX | 2.49e-36 | 918 | 1095 | 42 | 181 | Translin-associated factor-X (TRAX). TRAX (translin-associated factor-X) is a paralog of its binding partner protein Translin and together they form oligomeric complexes known as C3PO proteins (for component 3 promoter of RNA-induced silencing complex or RISC). TRAX complexed with Translin is possibly involved in dendritic RNA processing and in DNA double-strand break repair as an interacting partner with C1D, an activator of the DNA-dependent protein kinase involved in the repair of DNA-double strand breaks. It has been shown that Trax subunit, but not Translin, possesses a Glu-Glu-Asp catalytic center with the capacity to digest RNA; this catalytic activity is required for passenger-strand removal and RISC activation in RNAi. In Archaeoglobus fulgidus, Trax-like-subunits assemble into an octameric structure, highly similar to human C3PO; its complex with duplex RNA reveals that the octamer entirely encapsulates a single 13-base-pair RNA duplex inside a large inner cavity. Translin and Trax participate in a variety of nucleic acid metabolism pathways in addition to RNAi and have been implicated in a wide range of biological activities, including mRNA processing, cell growth regulation, spermatogenesis, neuronal development/function, genome stability regulation and carcinogenesis; however, their precise role in some of the processes remains unclear. |
| 271348 | Translin-like | 3.09e-16 | 914 | 1093 | 44 | 197 | Translin and translin-associated factor-X (TRAX). Translin (also known as TB-RBP), and its binding partner protein TRAX (translin-associated factor-X) are a paralogous pair of conserved proteins, and oligomeric complexes of TRAX and translin are known as C3PO proteins (for component 3 promoter of RNA-induced silencing complex or RISC). The Translin-Trax complex enhances the removal of the passenger strand in RNAi and the formation of active RISC. Translin and Trax participate in a variety of nucleic acid metabolism pathways in addition to RNAi and have been implicated in a wide range of biological activities, including mRNA processing, cell growth regulation, spermatogenesis, neuronal development/function, genome stability regulation and carcinogenesis; however, their precise role in some of the processes remains unclear. It has been shown that Trax subunit, but not Translin, possesses a Glu-Glu-Asp catalytic center with the capacity to digest RNA as well as DNA; this catalytic activity is required for passenger-strand removal and RISC activation in RNAi. In Archaeoglobus fulgidus, Trax-like-subunits assemble into an octameric structure, highly similar to human C3PO; its complex with duplex RNA reveals that the octamer entirely encapsulates a single 13-base-pair RNA duplex inside a large inner cavity. |
| 407658 | Mannosidase_ig | 1.29e-15 | 676 | 766 | 1 | 91 | Mannosidase Ig/CBM-like domain. This domain corresponds to domain 4 in the structure of Bacteroides thetaiotaomicron beta-mannosidase, BtMan2A. This domain has an Ig-like fold. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 808 | 1 | 809 | |
| 0.0 | 1 | 808 | 1 | 809 | |
| 0.0 | 1 | 808 | 1 | 810 | |
| 0.0 | 1 | 808 | 1 | 810 | |
| 0.0 | 1 | 808 | 2 | 810 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.80e-143 | 1 | 658 | 21 | 660 | Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12],5N6U_B Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12],5N6U_C Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12],5N6U_D Crystal structure of Beta-D-Mannosidase from Dictyoglomus thermophilum. [Dictyoglomus thermophilum H-6-12] |
|
| 6.42e-124 | 9 | 667 | 11 | 667 | Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VJX_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VL4_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VL4_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VMF_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VMF_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VO5_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VO5_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VOT_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VOT_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron VPI-5482],2VQT_A Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron],2VQT_B Structural and biochemical evidence for a boat-like transition state in beta-mannosidases [Bacteroides thetaiotaomicron],2VR4_A Transition-state mimicry in mannoside hydrolysis: characterisation of twenty six inhibitors and insight into binding from linear free energy relationships and 3-D structure [Bacteroides thetaiotaomicron VPI-5482],2VR4_B Transition-state mimicry in mannoside hydrolysis: characterisation of twenty six inhibitors and insight into binding from linear free energy relationships and 3-D structure [Bacteroides thetaiotaomicron VPI-5482] |
|
| 6.75e-124 | 9 | 667 | 13 | 669 | Structure of a beta-mannosidase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482],2JE8_B Structure of a beta-mannosidase from Bacteroides thetaiotaomicron [Bacteroides thetaiotaomicron VPI-5482] |
|
| 6.92e-124 | 9 | 667 | 13 | 669 | Chain A, Beta-mannosidase [Bacteroides thetaiotaomicron VPI-5482],7OP6_B Chain B, Beta-mannosidase [Bacteroides thetaiotaomicron VPI-5482],7OP7_A Chain A, Beta-mannosidase [Bacteroides thetaiotaomicron VPI-5482],7OP7_B Chain B, Beta-mannosidase [Bacteroides thetaiotaomicron VPI-5482] |
|
| 1.78e-123 | 9 | 667 | 11 | 667 | Structure of the Michaelis complex of beta-mannosidase, Man2A, provides insight into the conformational itinerary of mannoside hydrolysis [Bacteroides thetaiotaomicron VPI-5482],2WBK_B Structure of the Michaelis complex of beta-mannosidase, Man2A, provides insight into the conformational itinerary of mannoside hydrolysis [Bacteroides thetaiotaomicron VPI-5482] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 1 | 808 | 1 | 809 | Beta-mannosidase B OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=mndB PE=3 SV=3 |
|
| 0.0 | 1 | 808 | 1 | 810 | Beta-mannosidase B OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=mndB PE=3 SV=1 |
|
| 0.0 | 1 | 808 | 1 | 809 | Beta-mannosidase B OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=mndB PE=3 SV=1 |
|
| 0.0 | 1 | 808 | 1 | 810 | Beta-mannosidase B OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=mndB PE=3 SV=1 |
|
| 0.0 | 1 | 808 | 1 | 808 | Beta-mannosidase B OS=Aspergillus terreus (strain NIH 2624 / FGSC A1156) OX=341663 GN=mndB PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
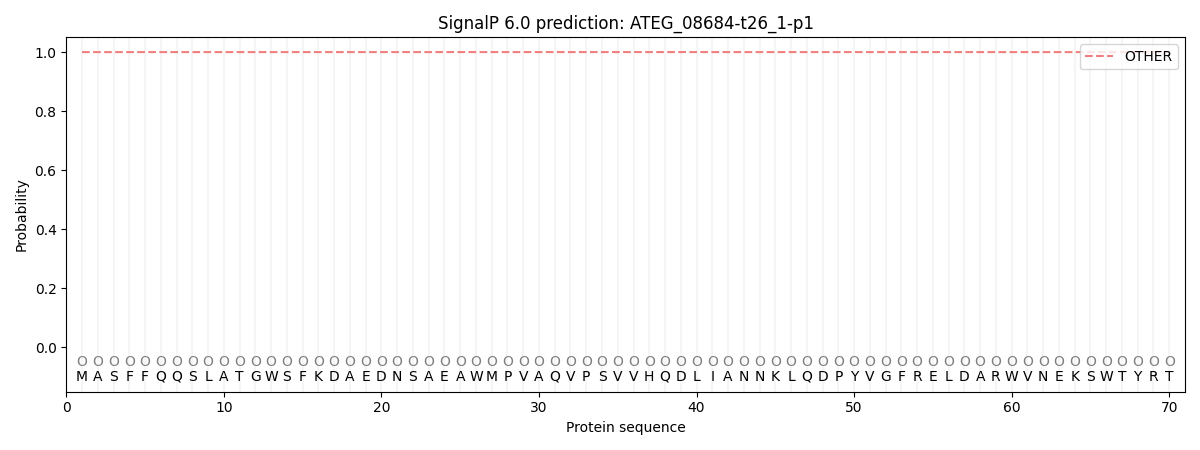
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000037 | 0.000000 |
