You are browsing environment: FUNGIDB
CAZyme Information: ATEG_08214-t26_1-p1
You are here: Home > Sequence: ATEG_08214-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus terreus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus terreus | |||||||||||
| CAZyme ID | ATEG_08214-t26_1-p1 | |||||||||||
| CAZy Family | GH92 | |||||||||||
| CAZyme Description | predicted protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA7 | 126 | 569 | 8.1e-53 | 0.9454148471615721 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396238 | FAD_binding_4 | 2.88e-18 | 125 | 268 | 5 | 136 | FAD binding domain. This family consists of various enzymes that use FAD as a co-factor, most of the enzymes are similar to oxygen oxidoreductase. One of the enzymes Vanillyl-alcohol oxidase (VAO) has a solved structure, the alignment includes the FAD binding site, called the PP-loop, between residues 99-110. The FAD molecule is covalently bound in the known structure, however the residue that links to the FAD is not in the alignment. VAO catalyzes the oxidation of a wide variety of substrates, ranging form aromatic amines to 4-alkylphenols. Other members of this family include D-lactate dehydrogenase, this enzyme catalyzes the conversion of D-lactate to pyruvate using FAD as a co-factor; mitomycin radical oxidase, this enzyme oxidizes the reduced form of mitomycins and is involved in mitomycin resistance. This family includes MurB an UDP-N-acetylenolpyruvoylglucosamine reductase enzyme EC:1.1.1.158. This enzyme is involved in the biosynthesis of peptidoglycan. |
| 223354 | GlcD | 2.84e-10 | 125 | 268 | 36 | 167 | FAD/FMN-containing dehydrogenase [Energy production and conversion]. |
| 369658 | BBE | 3.39e-09 | 531 | 568 | 1 | 38 | Berberine and berberine like. This domain is found in the berberine bridge and berberine bridge- like enzymes which are involved in the biosynthesis of numerous isoquinoline alkaloids. They catalyze the transformation of the N-methyl group of (S)-reticuline into the C-8 berberine bridge carbon of (S)-scoulerine. |
| 273751 | FAD_lactone_ox | 0.006 | 197 | 283 | 77 | 162 | sugar 1,4-lactone oxidases. This model represents a family of at least two different sugar 1,4 lactone oxidases, both involved in synthesizing ascorbic acid or a derivative. These include L-gulonolactone oxidase (EC 1.1.3.8) from rat and D-arabinono-1,4-lactone oxidase (EC 1.1.3.37) from Saccharomyces cerevisiae. Members are proposed to have the cofactor FAD covalently bound at a site specified by Prosite motif PS00862; OX2_COVAL_FAD; 1. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 1.40e-49 | 16 | 580 | 492 | 1055 | |
| 1.40e-49 | 16 | 580 | 492 | 1055 | |
| 3.41e-14 | 119 | 576 | 65 | 497 | |
| 2.21e-11 | 100 | 568 | 42 | 482 | |
| 1.19e-10 | 125 | 568 | 65 | 482 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.62e-78 | 23 | 590 | 27 | 574 | Crystal structure of VAO-type flavoprotein MtVAO615 at pH 7.5 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F73_A Crystal structure of VAO-type flavoprotein MtVAO615 at pH 5.0 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F73_B Crystal structure of VAO-type flavoprotein MtVAO615 at pH 5.0 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464] |
|
| 1.53e-56 | 11 | 586 | 17 | 597 | Crystal structure of VAO-type flavoprotein MtVAO713 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F74_B Crystal structure of VAO-type flavoprotein MtVAO713 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F74_C Crystal structure of VAO-type flavoprotein MtVAO713 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464],6F74_D Crystal structure of VAO-type flavoprotein MtVAO713 from Myceliophthora thermophila C1 [Thermothelomyces thermophilus ATCC 42464] |
|
| 1.41e-14 | 194 | 307 | 109 | 223 | Physcomitrella patens BBE-like 1 variant D396N [Physcomitrium patens],6EO5_B Physcomitrella patens BBE-like 1 variant D396N [Physcomitrium patens] |
|
| 1.41e-14 | 194 | 307 | 109 | 223 | Physcomitrella patens BBE-like 1 wild-type [Physcomitrium patens],6EO4_B Physcomitrella patens BBE-like 1 wild-type [Physcomitrium patens] |
|
| 1.91e-13 | 90 | 320 | 40 | 241 | Phl p 4 I153V N158H variant, a glucose oxidase [Phleum pratense],4PWC_A Phl p 4 I153V N158H variant, a glucose oxidase, 3.5 M NaBr soak [Phleum pratense] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 9.78e-173 | 24 | 586 | 26 | 587 | Uncharacterized FAD-linked oxidoreductase ARB_02478 OS=Arthroderma benhamiae (strain ATCC MYA-4681 / CBS 112371) OX=663331 GN=ARB_07056 PE=1 SV=2 |
|
| 1.35e-133 | 7 | 590 | 8 | 586 | FAD-linked oxidoreductase malF OS=Malbranchea aurantiaca OX=78605 GN=malF PE=1 SV=1 |
|
| 6.01e-122 | 22 | 587 | 29 | 581 | FAD-linked oxidoreductase notD' OS=Aspergillus versicolor OX=46472 GN=notD' PE=1 SV=1 |
|
| 2.03e-118 | 21 | 587 | 28 | 589 | FAD-linked oxidoreductase notD OS=Aspergillus sp. (strain MF297-2) OX=877550 GN=notD PE=1 SV=1 |
|
| 4.99e-115 | 23 | 585 | 26 | 581 | FAD-linked oxidoreductase easE OS=Epichloe festucae var. lolii OX=73839 GN=easE PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
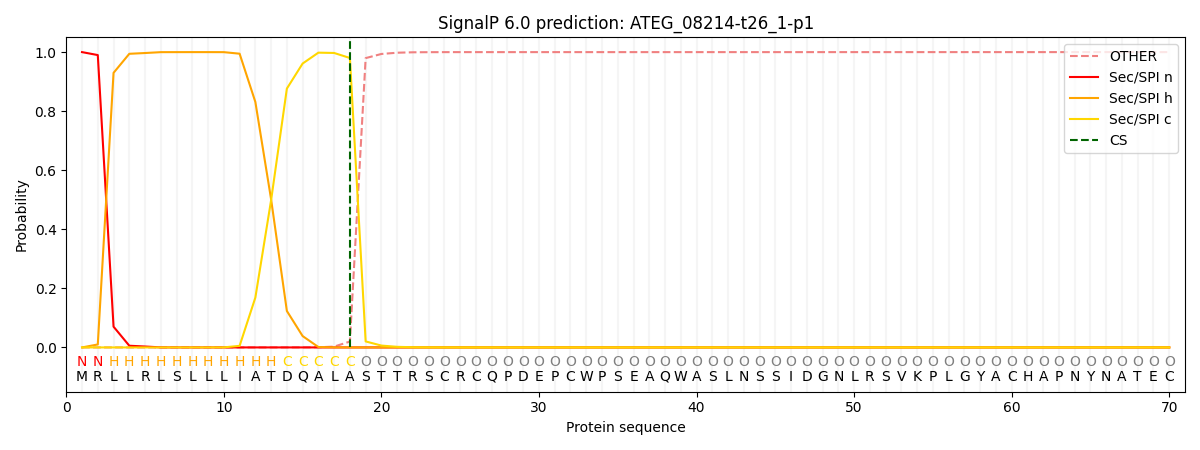
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000218 | 0.999740 | CS pos: 18-19. Pr: 0.9800 |
