You are browsing environment: FUNGIDB
CAZyme Information: ATEG_01675-t26_1-p1
You are here: Home > Sequence: ATEG_01675-t26_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus terreus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus terreus | |||||||||||
| CAZyme ID | ATEG_01675-t26_1-p1 | |||||||||||
| CAZy Family | AA9 | |||||||||||
| CAZyme Description | conserved hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| CE18 | 315 | 682 | 2.6e-189 | 0.997275204359673 |
| CBM87 | 97 | 314 | 6.4e-116 | 0.9954128440366973 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 396211 | Polysacc_deac_1 | 0.005 | 451 | 513 | 58 | 115 | Polysaccharide deacetylase. This domain is found in polysaccharide deacetylase. This family of polysaccharide deacetylases includes NodB (nodulation protein B from Rhizobium) which is a chitooligosaccharide deacetylase. It also includes chitin deacetylase from yeast, and endoxylanases which hydrolyzes glucosidic bonds in xylan. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 83 | 751 | 133 | 801 | |
| 0.0 | 83 | 751 | 133 | 801 | |
| 0.0 | 80 | 751 | 132 | 804 | |
| 0.0 | 1 | 751 | 1 | 794 | |
| 0.0 | 88 | 751 | 140 | 804 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 0.0 | 87 | 751 | 20 | 684 | Crystal structure of Agd3 a novel carbohydrate deacetylase [Aspergillus fumigatus Af293] |
Swiss-Prot Hits help
SignalP and Lipop Annotations help
This protein is predicted as SP
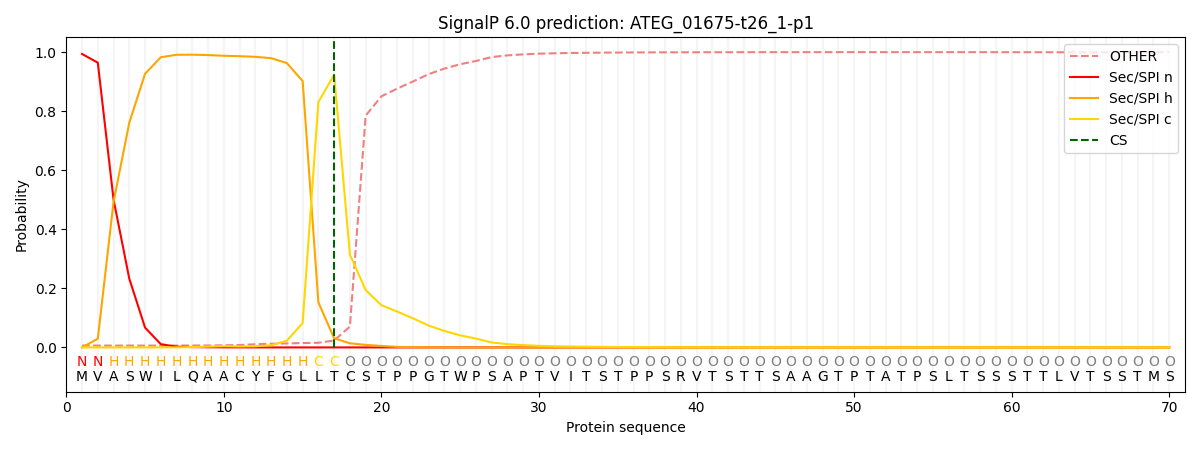
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.008412 | 0.991581 | CS pos: 17-18. Pr: 0.9237 |
