You are browsing environment: FUNGIDB
CAZyme Information: ATCC64974_96320-t41_1-p1
You are here: Home > Sequence: ATCC64974_96320-t41_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus niger | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus niger | |||||||||||
| CAZyme ID | ATCC64974_96320-t41_1-p1 | |||||||||||
| CAZy Family | GT34 | |||||||||||
| CAZyme Description | unspecified product | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
Enzyme Prediction help
| EC | 3.2.1.149:1 | 3.2.1.-:1 |
|---|
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH5 | 78 | 357 | 8e-156 | 0.9964285714285714 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 225344 | BglC | 3.52e-08 | 21 | 202 | 10 | 191 | Aryl-phospho-beta-D-glucosidase BglC, GH1 family [Carbohydrate transport and metabolism]. |
| 395098 | Cellulase | 7.42e-05 | 81 | 310 | 24 | 236 | Cellulase (glycosyl hydrolase family 5). |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 5.31e-230 | 1 | 382 | 1 | 327 | |
| 1.48e-186 | 15 | 384 | 20 | 389 | |
| 8.42e-179 | 12 | 384 | 16 | 388 | |
| 1.89e-178 | 20 | 375 | 29 | 383 | |
| 1.89e-178 | 20 | 375 | 29 | 383 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3.31e-180 | 20 | 375 | 10 | 364 | Chain A, Glycoside hydrolase family 5 [Aspergillus oryzae],6LA0_B Chain B, Glycoside hydrolase family 5 [Aspergillus oryzae] |
|
| 1.03e-161 | 19 | 371 | 12 | 365 | Crystal structure of rutinosidase from Aspergillus niger [Aspergillus niger],6I1A_B Crystal structure of rutinosidase from Aspergillus niger [Aspergillus niger] |
|
| 1.58e-11 | 31 | 316 | 5 | 302 | Chain A, Exo-beta-1,3-glucanase [uncultured bacterium],6ZB9_B Chain B, Exo-beta-1,3-glucanase [uncultured bacterium] |
|
| 8.93e-11 | 31 | 316 | 5 | 302 | Chain A, Exo-beta-1,3-glucanase variant E167Q/E295Q [uncultured bacterium],6ZB8_B Chain B, Exo-beta-1,3-glucanase variant E167Q/E295Q [uncultured bacterium] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.62e-25 | 31 | 363 | 44 | 353 | Probable glucan 1,3-beta-glucosidase A OS=Aspergillus niger (strain CBS 513.88 / FGSC A1513) OX=425011 GN=exgA PE=3 SV=1 |
|
| 9.38e-24 | 31 | 311 | 44 | 310 | Probable glucan 1,3-beta-glucosidase A OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=exgA PE=3 SV=1 |
|
| 8.21e-23 | 31 | 311 | 44 | 310 | Probable glucan 1,3-beta-glucosidase A OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=exgA PE=3 SV=1 |
|
| 1.12e-22 | 31 | 311 | 44 | 310 | Probable glucan 1,3-beta-glucosidase A OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=exgA PE=3 SV=1 |
|
| 3.80e-22 | 16 | 311 | 29 | 309 | Probable glucan 1,3-beta-glucosidase A OS=Aspergillus clavatus (strain ATCC 1007 / CBS 513.65 / DSM 816 / NCTC 3887 / NRRL 1 / QM 1276 / 107) OX=344612 GN=exgA PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as SP
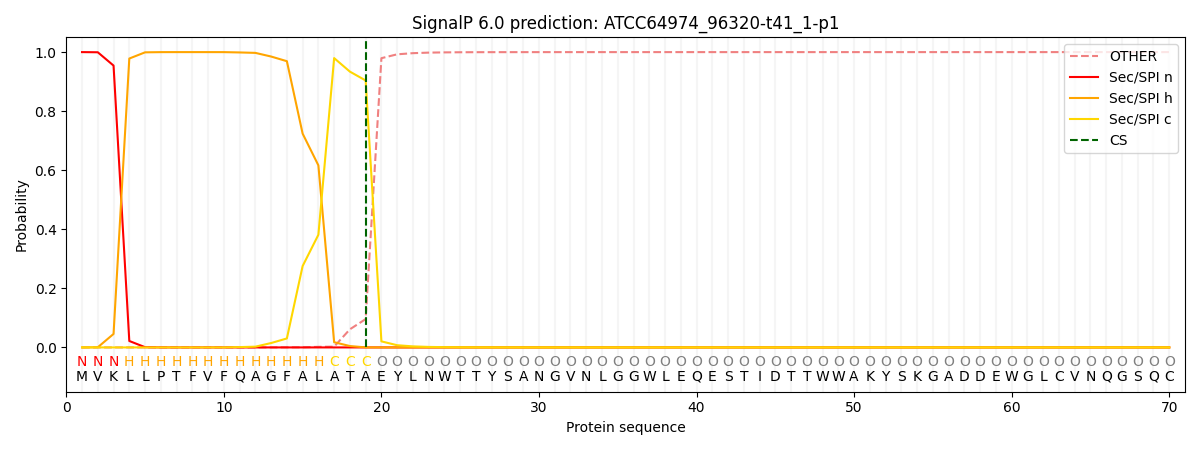
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 0.000238 | 0.999734 | CS pos: 19-20. Pr: 0.9039 |
