You are browsing environment: FUNGIDB
CAZyme Information: ASPZODRAFT_29543-t33_1-p1
You are here: Home > Sequence: ASPZODRAFT_29543-t33_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Penicilliopsis zonata | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Penicilliopsis; Penicilliopsis zonata | |||||||||||
| CAZyme ID | ASPZODRAFT_29543-t33_1-p1 | |||||||||||
| CAZy Family | GH55 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA9 | 12 | 194 | 8.5e-39 | 0.8227272727272728 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 410622 | LPMO_AA9 | 7.66e-36 | 14 | 211 | 20 | 216 | lytic polysaccharide monooxygenase (LPMO) auxiliary activity family 9 (AA9). AA9 proteins are copper-dependent lytic polysaccharide monooxygenases (LPMOs) involved in the cleavage of cellulose chains with oxidation of carbons C1 and/or C4 and C6. Activities include lytic cellulose monooxygenase (C1-hydroxylating) (EC 1.14.99.54) and lytic cellulose monooxygenase (C4-dehydrogenating) (EC 1.14.99.56). The family used to be called GH61 because weak endoglucanase activity had been demonstrated in some family members. |
| 397484 | Glyco_hydro_61 | 1.52e-26 | 16 | 164 | 27 | 174 | Glycosyl hydrolase family 61. Although weak endoglucanase activity has been demonstrated in several members of this family, they lack the clustered conserved catalytic acidic amino acids present in most glycoside hydrolases. Many members of this family lack measurable cellulase activity on their own, but enhance the activity of other cellulolytic enzymes. They are therefore unlikely to be true glycoside hydrolases. The subsrate-binding surface of this family is a flat Ig-like fold. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 2.15e-54 | 1 | 211 | 23 | 231 | |
| 1.21e-43 | 1 | 211 | 22 | 230 | |
| 4.03e-43 | 1 | 211 | 24 | 235 | |
| 4.93e-43 | 1 | 211 | 24 | 235 | |
| 6.94e-43 | 1 | 211 | 24 | 235 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.28e-18 | 54 | 211 | 56 | 212 | Lytic polysaccharide monooxygenase 9F from Neurospora crassa, NcLPMO9F [Neurospora crassa],4QI8_B Lytic polysaccharide monooxygenase 9F from Neurospora crassa, NcLPMO9F [Neurospora crassa] |
|
| 1.20e-16 | 51 | 211 | 49 | 206 | Zinc-bound glycoside hydrolase 61 E from Thielavia terrestris [Thermothielavioides terrestris],3EII_B Zinc-bound glycoside hydrolase 61 E from Thielavia terrestris [Thermothielavioides terrestris],3EII_C Zinc-bound glycoside hydrolase 61 E from Thielavia terrestris [Thermothielavioides terrestris],3EII_D Zinc-bound glycoside hydrolase 61 E from Thielavia terrestris [Thermothielavioides terrestris],3EJA_A Magnesium-bound glycoside hydrolase 61 isoform E from Thielavia terrestris [Thermothielavioides terrestris],3EJA_B Magnesium-bound glycoside hydrolase 61 isoform E from Thielavia terrestris [Thermothielavioides terrestris],3EJA_C Magnesium-bound glycoside hydrolase 61 isoform E from Thielavia terrestris [Thermothielavioides terrestris],3EJA_D Magnesium-bound glycoside hydrolase 61 isoform E from Thielavia terrestris [Thermothielavioides terrestris] |
|
| 2.73e-15 | 10 | 211 | 14 | 215 | Chain A, GLYCOSIDE HYDROLASE FAMILY 61 PROTEIN D [Phanerodontia chrysosporium],4B5Q_B Chain B, GLYCOSIDE HYDROLASE FAMILY 61 PROTEIN D [Phanerodontia chrysosporium] |
|
| 9.25e-13 | 74 | 211 | 78 | 225 | Crystal Structure of a Cellulose-active Polysaccharide Monooxygenase from M. thermophila (MtPMO3*) [Thermothelomyces thermophilus ATCC 42464],5UFV_B Crystal Structure of a Cellulose-active Polysaccharide Monooxygenase from M. thermophila (MtPMO3*) [Thermothelomyces thermophilus ATCC 42464],5UFV_C Crystal Structure of a Cellulose-active Polysaccharide Monooxygenase from M. thermophila (MtPMO3*) [Thermothelomyces thermophilus ATCC 42464],5UFV_D Crystal Structure of a Cellulose-active Polysaccharide Monooxygenase from M. thermophila (MtPMO3*) [Thermothelomyces thermophilus ATCC 42464],5UFV_E Crystal Structure of a Cellulose-active Polysaccharide Monooxygenase from M. thermophila (MtPMO3*) [Thermothelomyces thermophilus ATCC 42464],5UFV_F Crystal Structure of a Cellulose-active Polysaccharide Monooxygenase from M. thermophila (MtPMO3*) [Thermothelomyces thermophilus ATCC 42464] |
|
| 1.08e-12 | 18 | 211 | 31 | 223 | Crystal structure of HiLPMO9B [Heterobasidion irregulare TC 32-1],5NNS_B Crystal structure of HiLPMO9B [Heterobasidion irregulare TC 32-1] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2.33e-12 | 16 | 211 | 42 | 232 | Probable endo-beta-1,4-glucanase D OS=Neosartorya fischeri (strain ATCC 1020 / DSM 3700 / CBS 544.65 / FGSC A1164 / JCM 1740 / NRRL 181 / WB 181) OX=331117 GN=eglD PE=3 SV=1 |
|
| 2.50e-12 | 16 | 211 | 43 | 232 | Probable endo-beta-1,4-glucanase D OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=eglD PE=3 SV=1 |
|
| 2.50e-12 | 16 | 211 | 43 | 232 | Probable endo-beta-1,4-glucanase D OS=Aspergillus oryzae (strain ATCC 42149 / RIB 40) OX=510516 GN=eglD PE=3 SV=1 |
|
| 1.09e-11 | 16 | 211 | 42 | 232 | Probable endo-beta-1,4-glucanase D OS=Neosartorya fumigata (strain CEA10 / CBS 144.89 / FGSC A1163) OX=451804 GN=eglD PE=3 SV=1 |
|
| 1.09e-11 | 16 | 211 | 42 | 232 | Probable endo-beta-1,4-glucanase D OS=Neosartorya fumigata (strain ATCC MYA-4609 / Af293 / CBS 101355 / FGSC A1100) OX=330879 GN=eglD PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
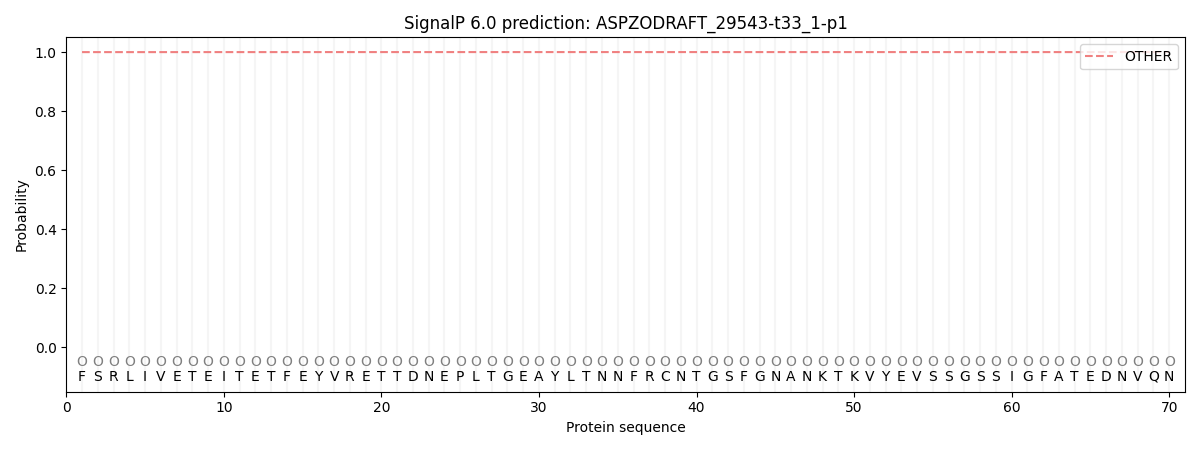
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000014 | 0.000048 |
