You are browsing environment: FUNGIDB
CAZyme Information: ASPTUDRAFT_59739-t33_1-p1
You are here: Home > Sequence: ASPTUDRAFT_59739-t33_1-p1
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Aspergillus tubingensis | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Ascomycota; Eurotiomycetes; ; Aspergillaceae; Aspergillus; Aspergillus tubingensis | |||||||||||
| CAZyme ID | ASPTUDRAFT_59739-t33_1-p1 | |||||||||||
| CAZy Family | GT20 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | ||||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| AA3 | 8 | 572 | 5.4e-156 | 0.9947183098591549 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| 235000 | PRK02106 | 1.29e-107 | 6 | 571 | 3 | 532 | choline dehydrogenase; Validated |
| 225186 | BetA | 3.50e-105 | 2 | 579 | 1 | 542 | Choline dehydrogenase or related flavoprotein [Lipid transport and metabolism, General function prediction only]. |
| 274888 | Rv0697 | 9.41e-62 | 10 | 570 | 2 | 486 | dehydrogenase, Rv0697 family. This model describes a set of dehydrogenases belonging to the glucose-methanol-choline oxidoreductase (GMC oxidoreductase) family. Members of the present family are restricted to Actinobacterial genome contexts containing also members of families TIGR03962 and TIGR03969 (the mycofactocin system), and are proposed to be uniform in function. |
| 398739 | GMC_oxred_C | 7.21e-45 | 427 | 565 | 1 | 142 | GMC oxidoreductase. This domain found associated with pfam00732. |
| 215420 | PLN02785 | 1.44e-28 | 8 | 563 | 55 | 569 | Protein HOTHEAD |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| 0.0 | 1 | 580 | 1 | 580 | |
| 0.0 | 1 | 580 | 1 | 580 | |
| 0.0 | 1 | 580 | 1 | 545 | |
| 3.88e-300 | 1 | 579 | 1 | 578 | |
| 5.34e-296 | 2 | 578 | 4 | 579 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 1.74e-261 | 9 | 578 | 7 | 577 | Crystal structure analysis of formate oxidase [Aspergillus oryzae RIB40],3Q9T_B Crystal structure analysis of formate oxidase [Aspergillus oryzae RIB40],3Q9T_C Crystal structure analysis of formate oxidase [Aspergillus oryzae RIB40] |
|
| 4.97e-261 | 9 | 578 | 7 | 577 | Effect of mutation (R554K) on FAD modification in Aspergillus oryzae RIB40formate oxidase [Aspergillus oryzae RIB40],5ZU3_B Effect of mutation (R554K) on FAD modification in Aspergillus oryzae RIB40formate oxidase [Aspergillus oryzae RIB40],5ZU3_C Effect of mutation (R554K) on FAD modification in Aspergillus oryzae RIB40formate oxidase [Aspergillus oryzae RIB40] |
|
| 1.42e-260 | 9 | 578 | 7 | 577 | Effect of mutation (R554A) on FAD modification in Aspergillus oryzae RIB40formate oxidase [Aspergillus oryzae RIB40],5ZU2_B Effect of mutation (R554A) on FAD modification in Aspergillus oryzae RIB40formate oxidase [Aspergillus oryzae RIB40],5ZU2_C Effect of mutation (R554A) on FAD modification in Aspergillus oryzae RIB40formate oxidase [Aspergillus oryzae RIB40] |
|
| 1.98e-74 | 9 | 572 | 2 | 563 | Crystal structure of aryl-alcohol oxidase from Pleurotus eryngii in complex with p-anisic acid [Pleurotus eryngii] |
|
| 1.08e-73 | 9 | 572 | 3 | 564 | Crystal structure of aryl-alcohol-oxidase from Pleurotus eryingii [Pleurotus eryngii] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 6.63e-72 | 4 | 573 | 2 | 536 | 4-pyridoxate dehydrogenase OS=Mesorhizobium japonicum (strain LMG 29417 / CECT 9101 / MAFF 303099) OX=266835 GN=padh1 PE=1 SV=1 |
|
| 2.73e-63 | 4 | 574 | 36 | 599 | Pyranose dehydrogenase 3 OS=Leucoagaricus meleagris OX=201219 GN=pdh3 PE=2 SV=1 |
|
| 8.74e-62 | 10 | 572 | 6 | 528 | Uncharacterized GMC-type oxidoreductase Mb1310 OS=Mycobacterium bovis (strain ATCC BAA-935 / AF2122/97) OX=233413 GN=BQ2027_MB1310 PE=3 SV=1 |
|
| 8.74e-62 | 10 | 572 | 6 | 528 | Uncharacterized GMC-type oxidoreductase MT1316 OS=Mycobacterium tuberculosis (strain CDC 1551 / Oshkosh) OX=83331 GN=MT1316 PE=3 SV=1 |
|
| 8.74e-62 | 10 | 572 | 6 | 528 | Uncharacterized GMC-type oxidoreductase Rv1279 OS=Mycobacterium tuberculosis (strain ATCC 25618 / H37Rv) OX=83332 GN=Rv1279 PE=1 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
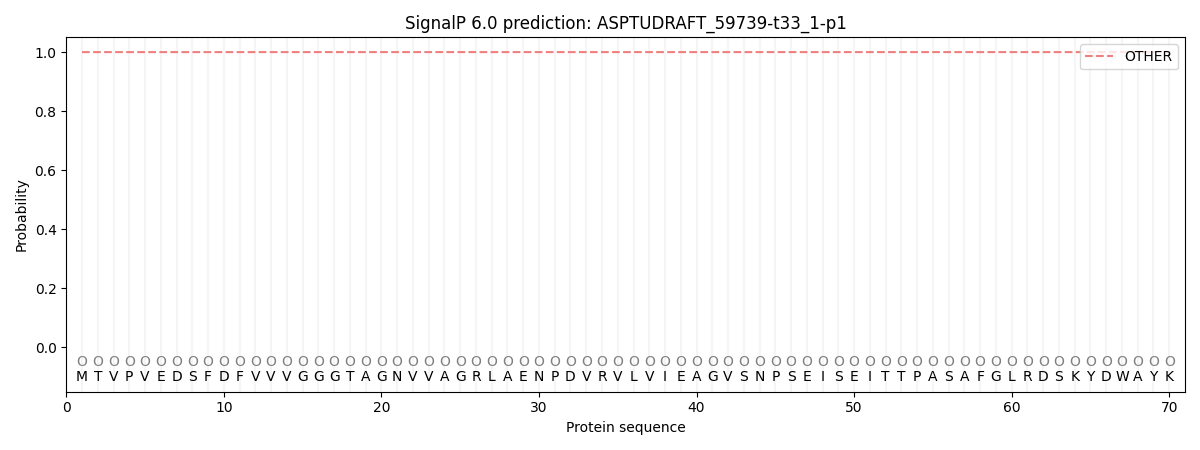
| Other | SP_Sec_SPI | CS Position |
|---|---|---|
| 1.000022 | 0.000011 |
